Metagenomic assembly algorithms
Developing tools for assembly of metagenomic sequence data.
The analysis of data from next generation sequencing of metagenomic samples has emerged as an important tool in recent years. In the past, much of this analysis has involved targeted 16S ribosomal sequencing followed by taxonomic classification. However, the increase in throughput and reduction in cost of NGS, combined with the lack of resolution provided by 16S approaches, has encouraged the adoption of whole genome shotgun approaches.
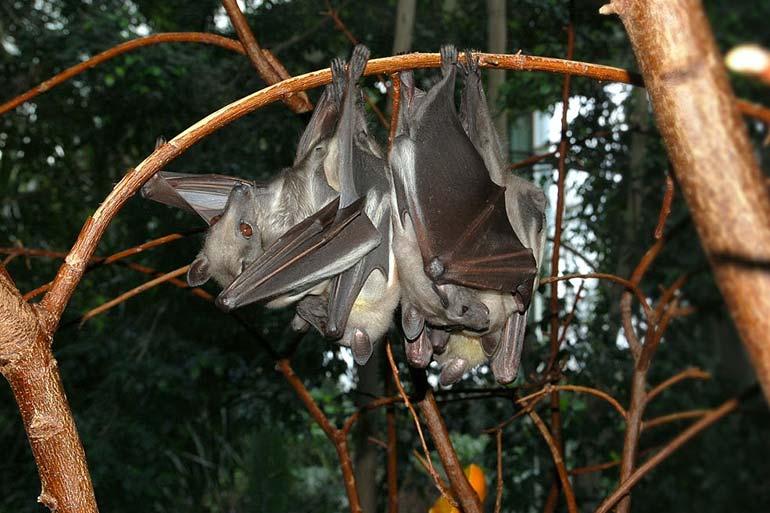
While read mapping is still a useful tool for analysing this data, greater insights are possible from assembly of reads. However, metagenomic assembly is still a relatively immature field with a handful of assemblers having emerged over the last few years. One of these is our own MetaCortex, a proof-of-concept assembly tool that has shown promising results when applied to the analysis of the virome of a species of bats from West Africa (Baker et al. 2013, Virology). The purpose of this project is to develop the algorithms necessary to turn the proof-of-concept into an efficient and sensitive assembly tool that will benefit the metagenomics community.
Straw-coloured fruit bat
Credit: By Fritz Geller-Grimm - Own work, CC BY-SA 2.5

Until recently, assembly approaches for metagenomic data have involved using standard de Bruijn graph assembly tools designed for single-organism genomic data. In a de Bruijn graph assembler, reads are broken down into overlapping k-mers, which form nodes in the graph. Nodes are linked together by edges that represent kmers that overlap in all but one base. Errors and repetitive kmers produce branches in the graph and the role of the assembler is to output contiguous sequence (contigs) by navigating paths through the graph. While such tools are capable of producing useful results, there are significant problems.
Genomic assembler heuristics tend to rely heavily on sequence coverage in order to simplify the graph and to find paths through it. This is a meaningful assumption in a standard genomic sample where the aim is to assemble a single genome from reads that can be assumed to be derived from the genome in relatively even coverage. However, in sequence reads from heterogeneous environmental samples, organisms tend to be represented at uneven levels of abundance, from partial genomes to high numbers of copies. Furthermore, common approaches taken by genomic assemblers to simplify the graph structure before building contigs – such as removal of tips and bubble structures – risk removing useful data from a metagenomic graph. The use of paired end information to resolve graph structure is also complicated in metagenomic data.
Such approaches rely on the use of read pairs to give support to particular paths through the graph at points of bifurcation; however in metagenomic datasets, there is much less likelihood that a single path through the graph will have strong enough support from paired end data. In our work, we are exploring alternative approaches to contig construction that do not rely on the traditional assumptions of genomic assemblers. We are also looking at approaches for data simplification in which we partition the readset in order to facilitate more accurate and longer contig construction. Finally, we are aiming to provide user-friendly tools that open up metagenomic assembly analysis to a wider audience.
MRC-University of Glasgow Centre for Virus Research
Techniques for assembly of metagenomic sequence data are in their infancy. As presented in the BBSRC's Review of Next Generation Sequencing, provision of assembly software for metagenomics is "highly deficient". An important academic impact of this work will be to drive forward methods for metagenomic assembly by increasing understanding of the problems, by developing new algorithmic approaches and by encouraging best practice techniques for analysis. The BBSRC's expert working group on metagenomics identified that the UK had failed to take full advantage of metagenomic techniques and this project will contribute to addressing this shortfall by helping to support the establishment of a research group focused on metagenomic tools and by increasing the knowledge and expertise of UK researchers.