Biography
I have been member of EI’s Core Bioinformatics Group since November 2017. I am tasked with building production-quality bioinformatics analysis pipelines and tools to add to the service portfolio of the group as well as running secondary analyses for customers. As a senior bioinformatician, I am also assisting with group managing and development. I am always curious to work on different bioinformatics (or computing) tasks and am happy to help, be it through consulting or training (e.g. new group members or students), or through establishing collaborations.
Background-wise, I am a “pure-blood” bioinformatician with a BSc (2004) and MSc (2006) from the Free University Berlin / Zuse Institute Berlin and a PhD (2011) from the Max Planck Institute of Molecular Plant Physiology (MPIMP) and University of Potsdam with a focus on RNA 3D structural bioinformatics. However, I have been working with genomics and transcriptomics data ever since my postdoc at the MPIMP (2011-2014), during which I also shifted from pure research to bioinformatics support, training, and tool/workflow development. Before joining EI (or, TGAC, as it was known back then) in January 2016 to work on wheat and grass genomics data, I briefly worked as Bioinformatics Support Officer at The Sainsbury Laboratory.
Projects
Publications
Related reading.

Genetic integrity needed for Biodiversity Net Gain to flower

Light-up plants and tunable roots signal new solutions for climate crisis

How the latest platforms are scaling-up our impact in aquaculture

The fish, the fungus, the grass, the bee - and the brassica

COPO: providing context through metadata

Standout innovation contributes to knowledge exchange
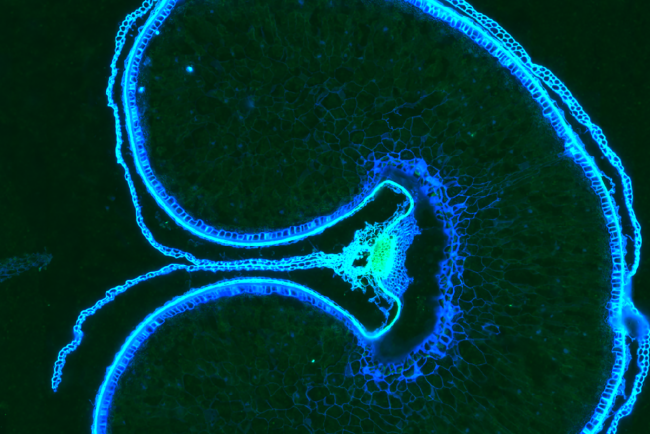
Applying spatial transcriptomics in plants

Collaborating for our future

Focus on fungi helps fight global threat to our food
New genome assembly finds yeast variant is distinct species

Science and Technology Secretary announces Engineering Biology investment

Identifying criminals from a single cell
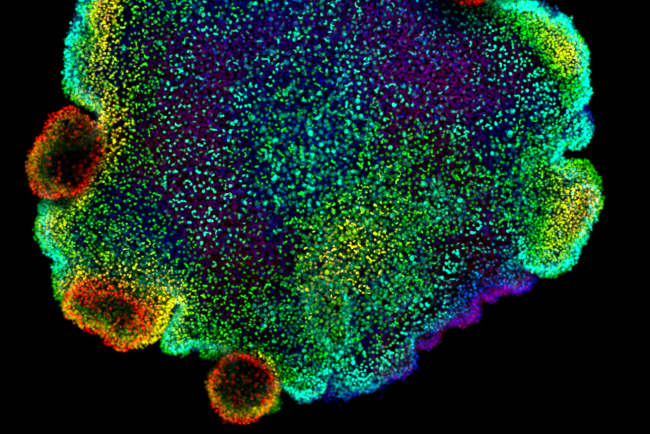
£3m funding for project to chart cellular diversity on Earth

Mysterious microbiomes to get makeover under transformational £5.4M grant
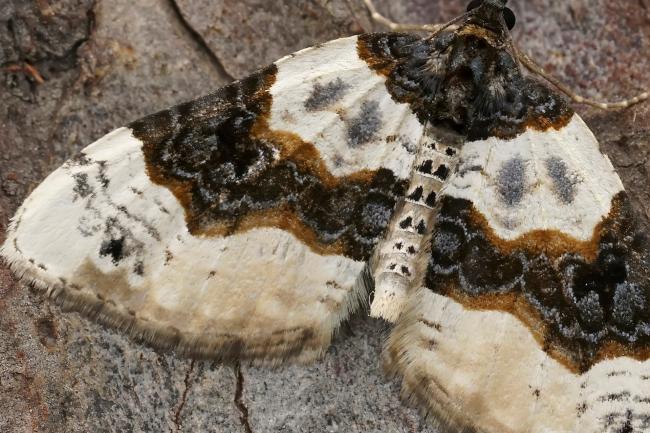

Purple Bar moth is 1,000th species sequenced in landmark project
Scientists one step closer to rewriting world’s first synthetic yeast genome




