Technical Genomics
Advancing genome science through technical leadership and innovation

Advancing genome science through technical leadership and innovation
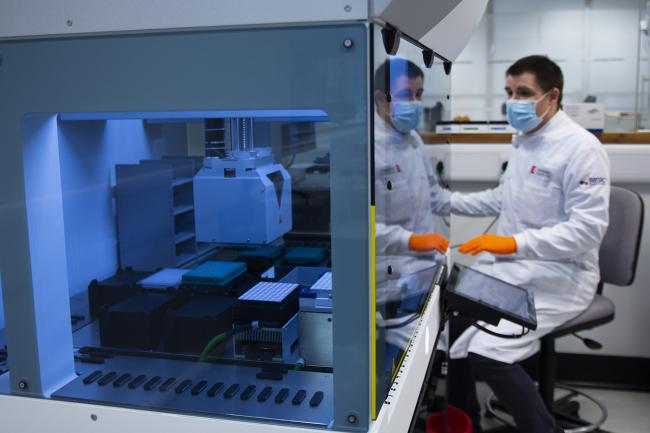
Technical Genomics, previously known as Genomics Pipelines, is a technology-focused group with specialist expertise in next-generation sequencing, laboratory automation, and information systems.
We develop, validate, and apply technical workflows to explore genomes, transcriptomes, and epigenomes across organisms and scale. Our work spans the full workflow cycle, from technical development to deployment in research projects and service delivery.
We make all our workflows accessible through the High-Performance Sequencing Platform, which we manage and operate in collaboration with the Core Bioinformatics group.
Our scientists work closely with the Technical Development and Core Bioinformatics groups to deliver the National Bioscience Research Infrastructure in Transformative Genomics, and collaborate with the wider Faculty to support the Institute Strategic Programmes.
While our technology of choice is DNA sequencing, our capabilities extend to a wider range of analytical techniques, from nucleic acid extraction to molecular diagnostics. We share our collective expertise with others through our participation in training activities and student placements.
Members of Technical Genomics are also actively involved in the life of the Institute, such as the Technician Commitment, Green Impact Programme, and IDEA Committee.
We are always open to collaboration and knowledge exchange. To work with us, please contact Karim Gharbi in the first instance.