Problems occur when the molecular machinery responsible for copying the DNA encounters an obstacle. This can cause things to slow down or briefly pause, which creates a narrow window of opportunity for mistakes to be made.
“The pausing of DNA replication leads to the accumulation of fragile, single-stranded DNA that is prone to base damage, slippage and double-strand breaks,” explains Nieduszynski. “Therefore, fork pausing is a major source of replicative errors, including point mutations, expansion/contraction of repeats, deletions and translocations.”
Although several checks and balances mean these obstacles normally get spotted and fixed during replication when things do go wrong the consequences can be catastrophic for the cell.
These rare but serious events are difficult to detect since most DNA replication is regular. Spotting them is something of a 'needle in a haystack' problem for researchers.
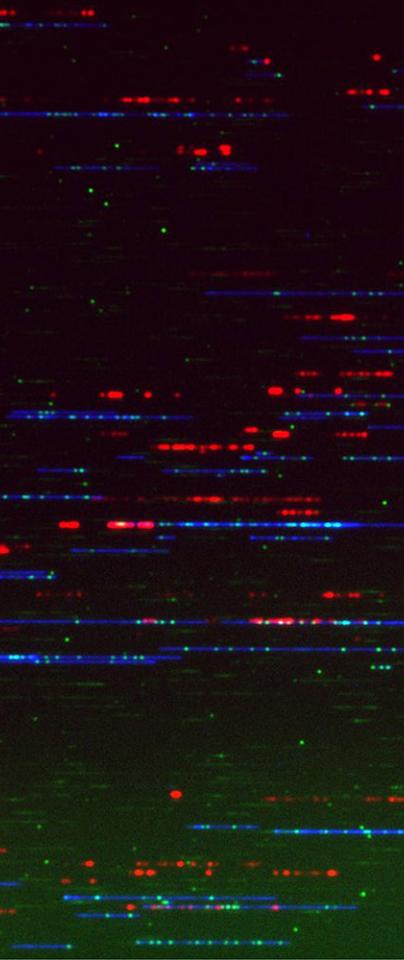
Nieduszynski’s Group has developed a high-throughput DNA sequencing technology that enables them to study the kinetics of DNA replication ‘in vivo’ on single molecules.
“This technology allows us to rapidly search for the 'needle in the haystack' and identify key molecular events, such as the slowing down or pausing of the DNA replication machinery,” he explains.
The Group is now applying this approach to determine what DNA sequences create obstacles to the DNA replication machinery, which protein factors assist in overcoming these hurdles, and how exactly pauses link to errors during the copying process.
“These approaches allow us to identify and characterise rare DNA replication mistakes, prioritising what determines and causes these mistakes and their resulting consequences,” says Nieduszynski.
“This is so important because a single DNA replication error on one chromosome in a single cell division has the potential to be harmless, lead to a subtle effect, or potentially be detrimental to the organism.”