Working with us
We welcome all enquiries from academia and industry. We would be happy to provide our expertise and facilities for the development of customized workflows or automation training.
Platform Lead:

The Earlham Biofoundry is strategically supported by BBSRC, part of UKRI, through the National Bioscience Research Infrastructure.
Access technical and scientific expertise
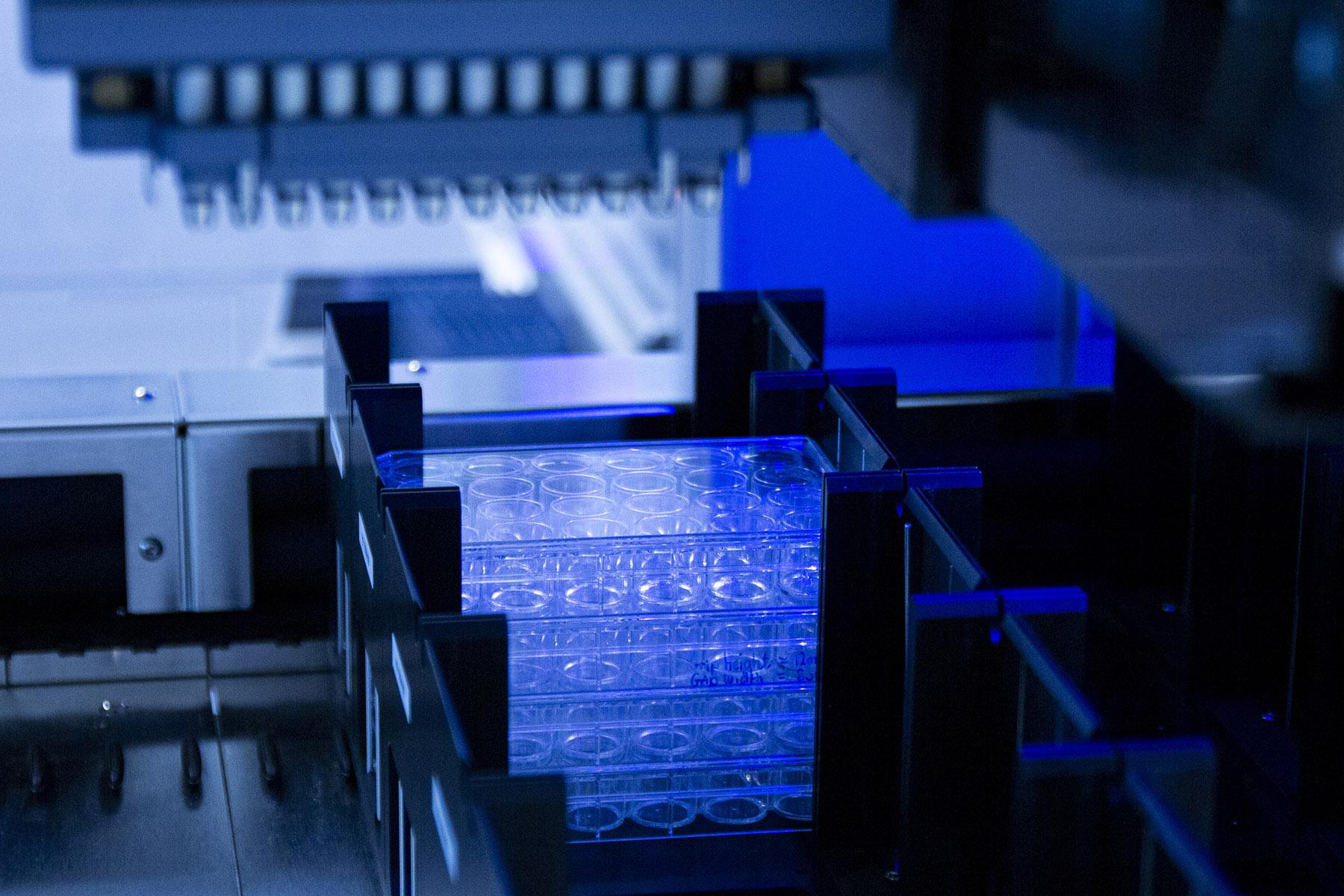
The Earlham Biofoundry is a resource for the UK biology and biotechnology communities, providing a platform to undertake large-scale projects.
We house suites of laboratory automation that, together with our expertise in large-scale experimentation and synthetic biology, can be applied to numerous molecular and microbiological workflows.
Synthetic Biology applies engineering principles such as standardisation and modularisation to biological sciences through the application of iterative design-build-test-learn cycles (DBTL). The standardisation of biological components and reactions allows experimental workflows to be automated.
This offers the potential to revolutionise the speed and scale of research, increasing accuracy and allowing miniaturisation to reduce costs. The adoption of these principles and approaches enables scientists to pursue complex experimental designs.
Established automated workflows are available to researchers across academia and industry and can be requested as a service. Alternatively, we can work collaboratively, contributing our scientific expertise and technical knowledge to develop new methods, automate, and scale up workflows, applying synthetic biology approaches to scientific problems.
We also provide training in laboratory automation through practical training sessions, laboratory placements, training courses and by hosting collaborators.
Service workflows
We offer access to a number of highly-optimised automated workflows on a fee-for-service basis. These are performed on our state-of-the-art automation platforms. We also make our automation methods and protocols available to the bioscience community on GitHub.
Nanoscale DNA Assembly, Including Combinatorial Libraries
Automated nanoscale DNA assembly of genetic parts and combinatorial DNA plasmid libraries using the Echo 650 platform. High-throughput construction of complex genetic designs, mostly using Golden Gate cloning or Gibson cloning.
Bacterial Transformation
Automated introduction of genetic constructs into bacterial hosts at scale using the Opentrons Flex, processing 8 to 96 samples per run. Integrated automated plating onto 6-well cyto plates using the Hamilton platform, eliminating manual spreading and ensuring reproducibility.
Colony Picking and Arraying
Automated high-throughput screening and arraying of E. coli colonies (blue/white and RFP/white selection) using the Hamilton STAR system, with capacity to pick up to 384 colonies in 2 hours.
DNA Purification
Automated DNA extractions using the CyBio Felix platform, processing 8 to 96 samples per run. Delivers DNA quantities suitable for pooling, sequencing, or downstream assemblies.
DNA Quantification and Normalisation
Automated DNA quantification and normalisation using the Opentrons Flex, ensuring accurate and consistent DNA concentrations across all samples. Available as a standalone service or integrated into larger automated workflows.
Automated High-Throughput DNA Electrophoresis (eGels)
Automated DNA electrophoresis using the Hamilton system, processing up to 96 using e-gels in 20 minutes.
High-Throughput Microfermentation
Miniaturised, high-throughput fermentation screening using the BioLector XT platform, enabling rapid evaluation of growth conditions, strain performance, and bioprocess parameters. Setup includes the LA Module for photosynthetic organisms and an anaerobic chamber, supporting a broad range of organisms. Ideal for proof-of-concept work to de-risk projects and optimise strains before scale-up. Actively seeking case study partners, get in touch if you are interested.
Cell-Free Protein Systems (CFPS) for Plants
Cell-free protein expression tailored for plant biology, enabling rapid prototyping of genetic constructs without the need for full plant transformation. Ideal for early-stage proof-of-concept work in agricultural biotechnology and plant-based biomanufacturing. For further reading, see our published paper.
Our automation capabilities

BioLector XT
The Biolector XT (Beckman Coulter) is a high-throughput microbioreactor (1-2 ml range) that enables real-time monitoring of key cultivation parameters, including biomass, fluorescence, pH, and dissolved oxygen. It supports both batch and fed-batch processes for aerobic, anaerobic, and phototrophic microorganisms, providing a versatile platform for efficient bioprocess development.

Opentrons Flex
Opentrons Flex
Opentrons are benchtop liquid handlers that are accessible and flexible enough to automate many common laboratory applications. They can be used to manipulate small volumes of liquids for undertaking molecular biology protocols, including PCR setup, serial dilutions, nucleic acid normalisation, and plate reformatting.
Visit the Opentrons Protocol Library for more information on applications.

Echo650 Beckman
Echo650 Beckman
Echo 650 is an Acoustic Liquid Handler ideal for high-throughput applications. It allows transfer of 2.5 nL solutions without the use of tips, minimising contamination risk and reagent waste. It enables assay miniaturisation across a broad range of applications, including Golden Gate assemblies in 2 µL total volume, Cell-Free Protein Synthesis (CFPS), and bacterial and yeast culture microdispensing into omnitrays.

Hamilton STAR and STARPlus
Hamilton STAR and STARPlus
Hamiltons are versatile liquid handling platforms with customisable deck layouts and a broad range of tools, including a labware gripper, pipetting channels, and an iSWAP Robotic Transport Arm. They offer superior measurement accuracy, anti-droplet control, and liquid level detection technologies. The platform has the versatility to work with 96-well and 384-well plates, utilising both 8-channel and 96-channel pipetting heads. In our foundry, we have established protocols for bacterial spreading in 6-well cytoplasmic plates, as well as automated colony picking (approximately 400 colonies in 3 hours) with integrated imaging to detect colonies by colorimetric output.

Pioreactor
Pioreactors provide a small-scale (15 m) fermentation system featuring compact, affordable, and user-friendly hardware, paired with open-source software. The system supports customizable and versatile applications, enabled by basic automated liquid handling and a pumping system. It allows for growth monitoring of yeast, bacteria, and algae, with features like temperature control and LED lighting for cultivating photosynthetic microorganisms, effectively functioning as photobioreactors. Additionally, Pioreactors enable the creation of clusters to control multiple units in parallel.

CyBioFelix
CyBioFelix
Felix platforms provide flexible and fully automated multi-channel pipetting on a unique two-level deck system with easy-to-change pipetting heads. In our Biofoundry, we have established an automated protocol for plasmid DNA extraction of up to 96 samples per run using magnetic bead purification kits.

CLARIOstar Plus
CLARIOstar Plus
Flexible Plate reader that allows plate-based detection of Fluorescence intensity, Luminescence and UV/vis absorbance spectra

Latest research
Engineering Synthetic Plant Chromosomes
We specialise in the rapid design, construction, and testing of genetically reprogrammed organisms, working with a range of different partners in academia and industry.
Funded by a multimillion pound grant from the UK Government’s Advanced Research + Invention Agency (ARIA) as part of the Synthetic Plants programme, a multidisciplinary collaboration of researchers is leveraging these advances to establish synthetic plant chromosome (synPAC) technologies.
As part of the project, scientists in the Earlham Biofoundry are assembling and testing a potato regulatory element library, using automated high-throughput DNA assembly workflows and protoplasts testing.