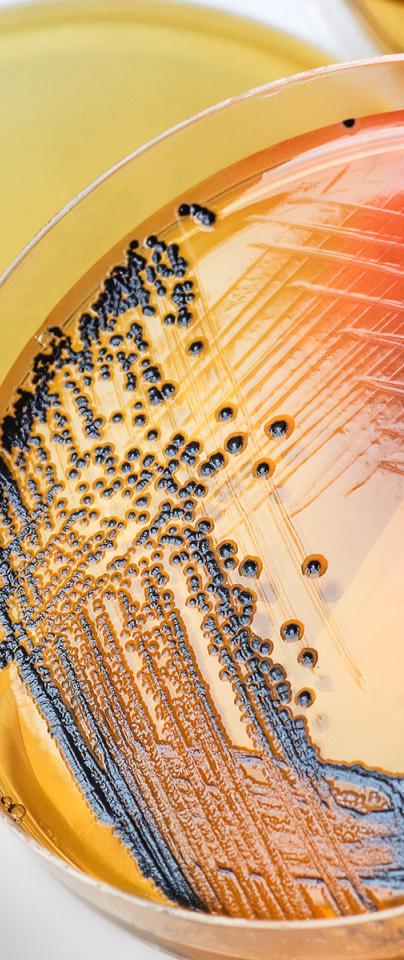
In parallel to the 10,000 Salmonella Genomes project, Professor Hall has led research focusing on how antibiotics in the environment affect the emergence of AMR in Salmonella. A brand-new sequencing approach for bacterial genomes was developed by the Earlham Institute specifically to study this.
Previously, genetic changes in bacteria were analysed either by bulk sequencing - taking an average from a very large quantity of cells - or laboriously isolating and characterising individual bacteria.
“There is a good analogy: if your head is on fire and your feet are in a freezer then, on average, you are fine,” says Professor Hall. “Averages do not give a high-resolution picture.
“And we have almost the opposite issue when we isolate a single cell - we have a clear picture of one bacterium, but we don’t know how representative that bacterium is. We could be seeing 90 per cent of the billions of cells in the culture, or one per cent.”
What was needed was the clarity of analysis from isolation combined with the large amount of data from a bulk culture. In other words, a system which was capable of producing a large quantity of single genomes at very high resolution.
This system – high-throughput single cell sequencing of whole genomes – has now been developed by the Institute and offers an unparalleled insight into evolution and diversity of Salmonella.
It gives crucial information about drug resistance and virulence factors - data which can be used to inform public health control strategies across the world.