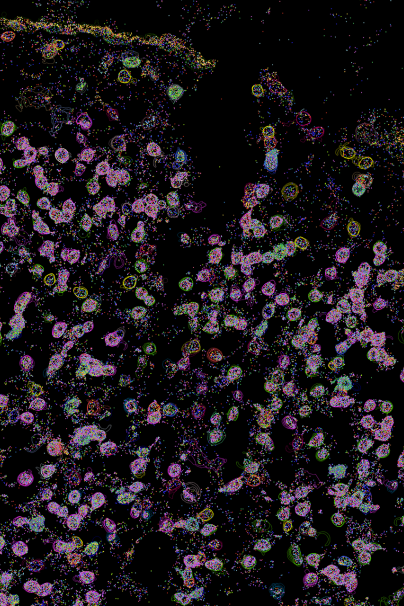
The afternoon session was led by Professor Monk.
He said: “I was really surprised at just how quick the area of single-cell biology is advancing, mostly thanks to innovative microfluidic and molecular technologies.
“In just a few years single-cell sequencing has evolved from processing a handful of cells to millions simultaneously. It is important that such momentum is maintained, especially with regard to medical disciplines.”
He described hearing numerous times during the symposium that unsupervised analyses of cell types and states allow for improved understanding of disease progression, providing new insights into clinical outcomes and patient stratification.
Adding: “I really look forward to seeing if some of the jaw-dropping methods, such as repeated cytoplasmic biopsy and single-cell multiple histone profiling, can be routinely incorporated into single-cell profiling workflows.”
In this session, Professor Charles ffrench-Constant, Pro-Vice-Chancellor for Norwich Medical School at UEA, presented his work using single nucleus RNA sequencing to explore patient heterogeneity in progressive multiple sclerosis.
Dr Rasa Elmentaite from the Wellcome Sanger Institute gave an overview of her PhD work creating single-cell atlases of the human gut, prioritising cells involved in disease.
The heterogeneity and sub-clonal diversity of tumour cells associated with resistance and metastasis in ER+ breast cancer was presented by Dr Shefali Thakur from the Institute of Cancer Research.
She was followed by Dr Leonie Luginbuehl from the University of Cambridge, who presented on a single cell gene expression atlas of rice and sorghum shoots during photomorphogenesis, and Dr Hayley Bennett, from Genentech, who spoke on increasing throughput and sequencing flexibility in single-cell multiomics workflows.
Dr Claudia Buhigas, from UEA, presented on single cell multi-omics profiles and how defective genomic activation and epigenetic reprogramming affects arrest in human pre-implantation embryos.
And Dr Andrew Bell from the University of Glasgow presented data, generated in collaboration with the Earlham Institute, which he is using to define the organisational logic of the anterolateral system in mice.