Biography
Falk is a Group Leader working across both the Earlham Institute and the Quadram Institute.
Falk is a bioinformatician with a passion for microbial ecosystems, bacterial evolution and developing computational systems to tackle these subjects in a combined perspective. Falk joined Earlham Institute in early 2019, developing metagenomic tools for tracking bacterial strains at high resolutions, to predict their genomic capabilities and explore their associations to diseases.
During his early career, Falk was working at the University of Constance (Germany) and the University of Sussex (UK) on bacterial evolution and tracking outbreaks of pathogens through following changes in their genomes. Moving from a genome centric view to a metagenomic view of whole microbial ecosystems, he worked during his PhD at the University of Brussels (Belgium) on bacterial associations to complex diseases such as IBD, obesity and Diabetes. For this, Falk developed bioinformatic pipelines to process 16S data (LotuS) as well as the statistical tools.
During his postdoc at EMBL Heidelberg (Germany), Falk continued his research on association to the human microbiome of complex diseases such as Diabetes and Parkinson’s disease, among others. His interests soon developed to track strains in the metagenomic datasets, as well as de novo assembling genomes from metagenomes. These techniques are mostly applied to the gut microbiome of mammals. Another interest of his is environmental metagenomics. For example, in 2018 he proved that in soils more fungi are usually connected to more antibiotic resistance in bacteria, and this is true on a local but also global scale.
Developing specialised algorithms that help answer complex questions on large biological datasets is the key ingredient for Falk’s scientific work.
Projects
Publications
Related reading.
Automation, engineering and computational biology: My Year in Industry
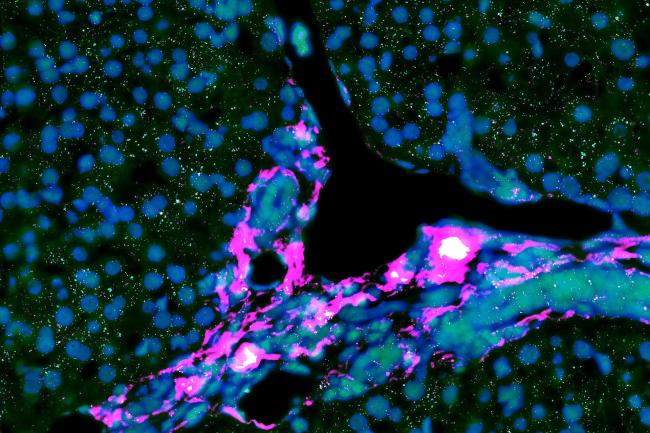
Advancing understanding of rare liver diseases through clinical and genomic collaboration

Breaking the silos: How open collaboration is transforming plant pangenomics

Seven ways Earlham Institute’s research improves future health

I told myself science wasn’t for me. I was wrong.

How single-cell genomics is unmasking the hidden diversity of protists

Forging a career as a research technical specialist

Decoding microbial diversity and function in irreplaceable habitats
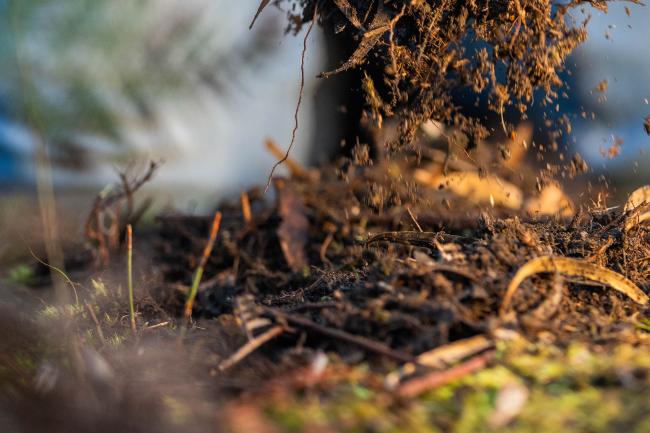

Metagenomic software advance boosts research into microbial diversity
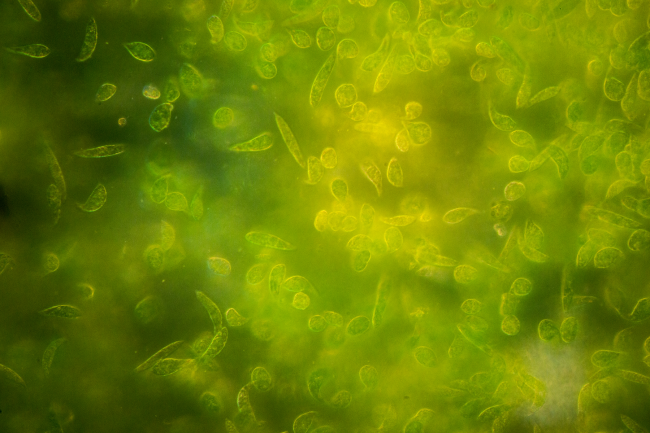
Unlocking the value of biodiversity in the UK and Ireland
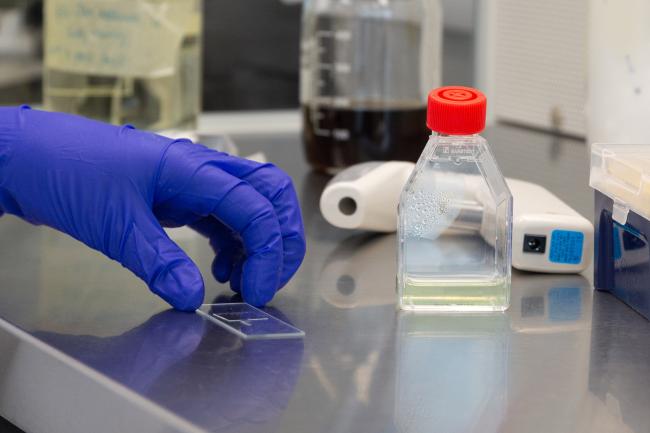

Single-cell sequencing reveals unexpected protist diversity
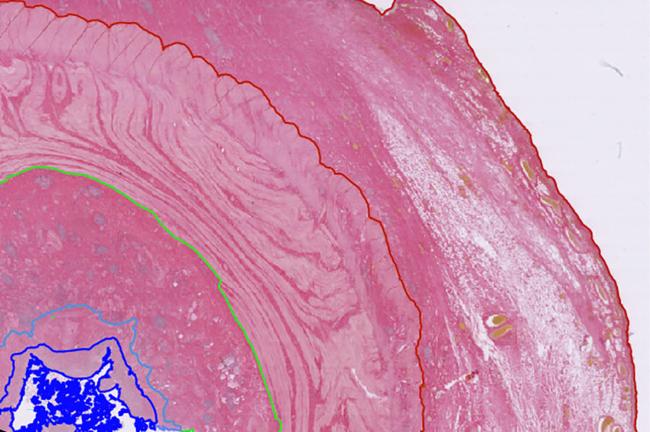
New study maps cellular mechanisms driving fibrosis in Crohn's Disease
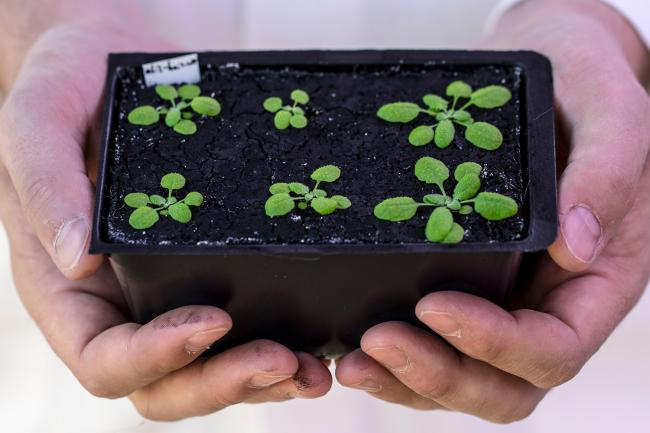

Maternal wisdom – how mother plants prime their seeds for success
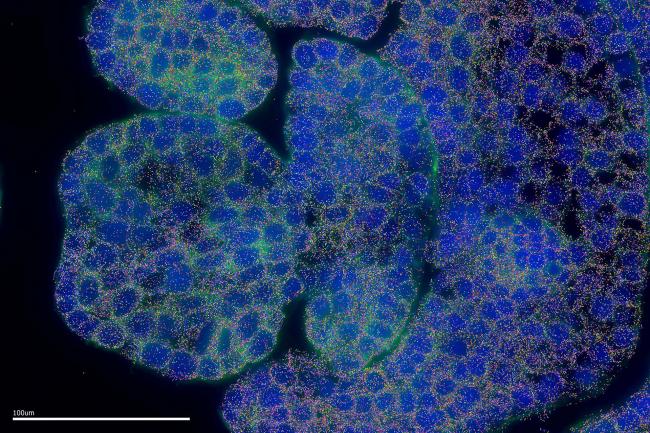
Innovations in spatial imaging could unlock higher wheat yields

Advancing forensic science: could a single cell prevent a wrongful conviction?

Earlham Institute PhD graduate recognised for excellence in scientific research