Around 500,000 people in the UK live with Inflammatory Bowel Disease (IBD), a life-long, chronic condition characterised by sporadic bouts of gut inflammation causing debilitating symptoms. Crohn’s Disease and Ulcerative Colitis – the latter affecting around 1 in 400 people - are the two most common types of IBD. Current treatments are ineffective and seriously impact the quality of life of the patients and those of their families.
The human gut harbours a community of microbes, known collectively as the microbiome, which is crucial to maintaining good health. A disrupted microbiome can cause gut-related conditions including IBD, an immune-linked inflammatory disease that causes abdominal pain, diarrhoea and extreme fatigue.
People with IBD tend to have reduced diversity or a change in the balance in their gut microbiome, especially of Bacteroides and Firmicutes bacteria. However, we still don’t know how exactly this translates to the triggering and progression of IBD. By understanding how these bacteria interact with the gut lining, and the immune system, and how this differs in IBD, we can better understand the causes and start developing targeted, effective treatments.
But to decipher this crosstalk across the different kingdoms of life, you need to understand how bacteria communicate, and then how human cells react to that information. This quest united microbiologist and immunologist Professor Simon Carding from the Quadram Institute and UEA, with Dr Tamás Korcsmáros, a systems biologist whose expertise lies in cellular signalling networks from the Earlham and the Quadram Institutes.
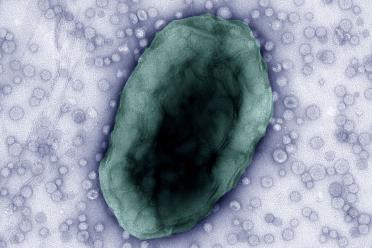
Professor Carding and his team have been investigating Bacterial Extracellular Vesicles (BEVs), which are tiny packages created by bacteria that they fill with various molecules and release from the cell. They can cross the gut lining, reaching cells of the immune system where they are recognised by receptors. The contents of the BEVs are molecular signals that then trigger the immune cells to react, with that signal potentially cascading into widespread effects.
In a healthy gut, BEVs and their cargo can contribute to anti-inflammatory responses of the immune system, but in an inflamed IBD patient’s gut, this response is lost. BEVs could therefore be used as a potential new therapy. But currently, we don’t understand enough about how they interact with the complex immune system. Our immune response relies on different types of cells monitoring for a plethora of different signals and interacting with each other to respond appropriately to a perceived threat of infection locally, and systematically across the body.
To address this knowledge gap, Dr Tamás Korcsmáros and his team used a previously published dataset about which genes are actively making proteins in 51 types of colon cells, from either healthy conditions or under the effect of ulcerative colitis. Uniquely, this dataset contained inflamed and uninflamed data from the same patients, allowing investigation of the effect of inflammation and not only the complex disease.
The team also analysed and characterised all of the cargo proteins obtained from BEVs made by the common gut bacterium Bacteroides thetaiotaomicron (Bt).
They then combined these datasets using an experimentally verified computational pipeline (called MicrobioLink) that predicts the interactions between microbial and host proteins, and how these triggers complex networks of cascading signalling systems. From this they could build up an overall picture of which microbial proteins were able to interact with which human proteins in the different types of immune cell and identify the differences between these networks in a healthy gut and in IBD.
This model, called an interactome, provides a snapshot of the constant communication between gut bacteria and our own immune system. From this the researchers could get an idea of the biological processes affected by microbial proteins - in healthy and inflamed UC conditions.
Their findings were published recently in the prestigious Journal of Extracellular Vesicles, one of the leading journals covering vesicle-mediated biological communication. The study was funded by the Biotechnology and Biological Sciences Research Council, part of UK Research and Innovation (UKRI).
Many interactions were identified common across cell types, but the research uncovered many biological processes that were specific to one type of immune cell. Focussing specifically on one pathway known to be important in immunity and inflammation, they were also able to identify differences between the same cell types in healthy and ulcerative colitis conditions. Experiments using cell cultures grown together with BEVs validated the predictions from the computational modelling.
“The finding that BEVs affect the immune system’s pathways in a cell-type specific manner, and that they are altered in inflammatory bowel disease is an important step to understanding the condition, and potentially could help in developing BEVs as a therapeutic system” said Lejla Gul, first author on the paper and an iCASE PhD student at the Earlham Institute and the Quadram Institute, supported by the BBSRC Norwich Research Park Biosciences Doctoral Training Partnership.
“Studying interkingdom connections with BEVs in a cell-type specific resolution requires multi-disciplinary expertise and various ‘omics datasets. Then you need a computational pipeline to analyse the data from different patients. Besides the actual scientific results, in the paper we introduce an open-source pipeline that others can use to analyse their data" said Dr Tamás Korcsmáros. “We hope that what we have demonstrated here in this study will be applied by others for understanding the mechanisms how other bacterial species communicate with our cells, and how it may be altered in other diseases.”
“This study highlights the importance and impact of laboratory scientists working with bioinformaticians to develop the means and tools for understanding the highly complex nature of the interactions between our gut microbes and cells of our body that is central to maintaining our health” said Professor Simon Carding. “The insights gained from studies such as this will be invaluable in developing new interventions aimed at maintaining health by promoting beneficial interactions with gut microbes and preventing harmful ones that can lead to diseases such as IBD.”
"Extracellular vesicles produced by the human commensal gut bacterium Bacteroides thetaiotaomicron affect host immune pathways in a cell-type specific manner that are altered in inflammatory bowel disease" is published in the Journal of Extracellular Vesicles.
Above image: Bacteroides thetaiotaomicron cell releasing Bacterial Extracellular Vesicles (BEVs), which are tiny packages created by bacteria that they fill with various molecules and release from the cell. Image by Rokas Juodeikis (Quadram Institute) and Ian Brown (University of Kent)