In terms of public impact, single-cell genomics techniques can be applied in pre-implantation genetic diagnosis, which allows screening of IVF embryos to detect genetic problems before implantation.
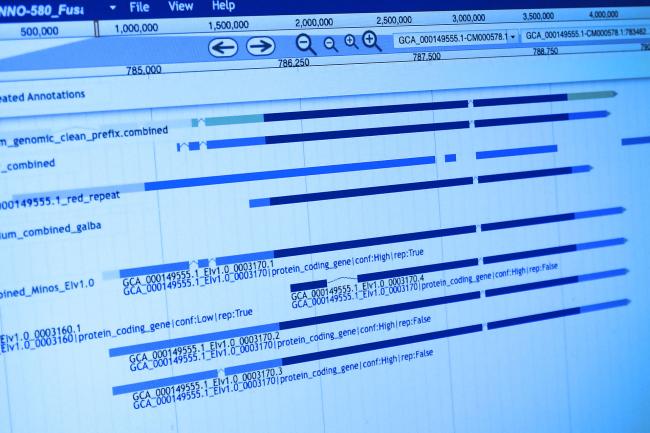
There is also a lot of work ongoing in cancer biology to use single-cell genomics to track the life history of a tumour – mutations unique to subsets of the cells within a tumour can be used to reconstruct a ‘lineage tree’ of the cancer cells. By knowing how the cells are related, and the ways in which they are mutated, it might be possible for clinicians to offer enhanced personalised treatments.
Beyond the immediately practical, single-cell genomics has the potential to reveal some of the true complexity of living things – there are more cells in one human body than there are stars in the Milky Way, many of them doing very different jobs and all of them arising from one single-cell after fertilisation.
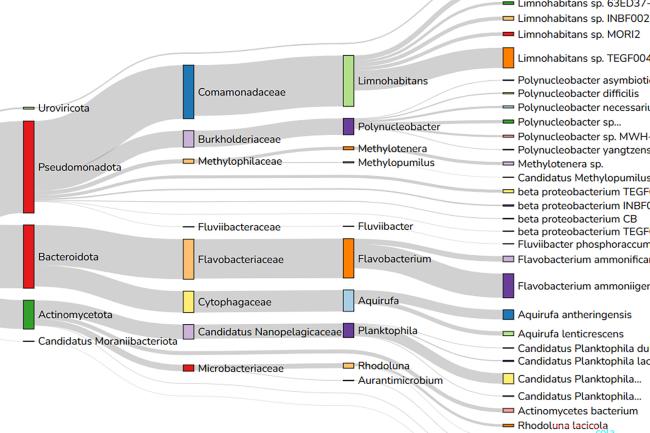
Even the populations of bacteria in our guts, or the cells in the root of a plant, are enormously diverse and complex. At the single-cell level, pretty much everything is amazing – and I’d like to think that by capturing some of this complexity and visualising it, single-cell genomics might capture some of the public’s imagination too.