Systems Genomics approaches to understand complex phenotypes
This programme seeks to identify and understand the functional roles of alleles in biological systems.

This research programme has now concluded. Please visit Our Research Programmes to find out about our current research.
The analysis and interpretation of genetic variation in genomes of organisms relevant to food security, enabled by algorithm and software development for accurate genome assembly and identification of sequence variants.
Our programme aims to advance our understanding of the impacts of selection and domestication at the genome level in economically important crop and aquaculture species. We also aim to further our understanding of foodborne pathogen virulence including antibiotic resistance evolution.
The delivery of these objectives requires significant algorithmic and software developments to enable the accurate assembly of genomes and the identification of genetic variation in both elite strains and wild relatives. We will also pursue development of novel algorithms for microbe identification and quantification including applications for real time and in-field monitoring.
The identification of regions and genes under natural or artificial selection in crops and aquatic species will enable breeders to identify genes that are important for productivity (yield, nutritive quality), accelerating breeding processes. The analysis of wild relatives adapted to challenging environments (pathogens, drought, cold) will lead to the identification of genomic regions and genes that could be exploited for the production of elite strains better adapted to environmental variation, and emerging threats. We will also deliver high-power datasets to breeders and other large scale crop and aquaculture research projects using novel infrastructure that will enable generation of better markers for breeding programmes.
Polyploidy has long been recognised as one of the hallmarks associated with domestication. Among the different mechanisms leading to ploidy variation, hybridization can result in new phenotypes not present in the parental strains allowing the exploitation of novel niches and environments. We will explore the impact of hybridization and genome duplication on gene regulation, including epigenetic marks and the evolution of regulatory elements. This will provide a better understanding of how genomes respond to introgression and other genomic diversification events. This fundamental knowledge will aid current efforts to generate diversity and improvement through the use of synthetic lines.
Our work on foodborne pathogens will be of interest to clinical researchers as well as the food industry both through the the characterisation of species and strains that cause disease, but also the relative pathogenicity associated with different bacterial strains. Our work on antimicrobial resistance will be of particular interest for the community to understand the implications of antimicrobial resistance as an adaptive trait and the mechanisms that can help prevent this.
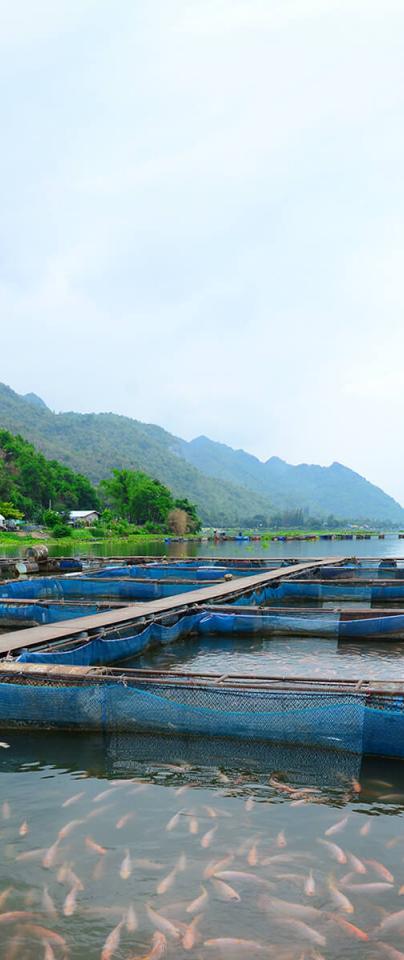

With increasing pressure from pests, disease, urbanisation and climate change, global food security must be kept robust if we are to feed a population expected to reach ten billion by 2050. Exploring the genetic signatures of domestication and adaptation is vital in supporting agricultural and aquacultural systems in a move towards sustainability.
Our work will help expand breeding programmes of farmed plants and animals, provide the data that breeders need to improve productivity, and uncover insights into antimicrobial resistance.

This programme seeks to identify and understand the functional roles of alleles in biological systems.
Data science for integrative biology

Coordinating expertise across BBSRC institutes with complementary university programmes to make sure researchers are well equipped to support the development of this crucial food crop.