10 years of impact: keeping living things healthy
From <i>Salmonella</i> to ash dieback: ten years of impact on how we understand human health and that of the environment.
From Salmonella to ash dieback: we’ve learned a lot about the key factors involved in the outbreak of disease and the evolution of pathogens and how we can use this knowledge to protect human health and that of the environment.
The interactions between all of the organisms in each of the living systems on Earth are complex, but at EI we’re determined to decode them for the benefit of us all. Some of these interactions, when thrown out of kilter, can lead to devastating infections of humans and plants alike.
From the devastating ash dieback disease that is rampaging through Britain’s trees to the worldwide menace that is Salmonella, how have researchers at EI been applying genomics and bioinformatics to better understand our health and that of our valued ecosystems - and what’s the value to the UK economy and beyond over the next 25 years?
We commissioned a report (Brookdale Consulting) on the impact of some highlights of our research so far.

A deadly disease to rival malaria, Salmonella, as featured in our recent article, is about more than just bad eggs. In recent times, worrying strains have caused alarmingly high mortality in immunocompromised patients in sub-Saharan Africa, while antibiotic resistant strains of the pathogen are spreading throughout the globe.
Human non-typhoidal Salmonella infections can be divided into two types: those restricted to the gut and those that reach the bloodstream which are known as invasive. Globally it is estimated that 93m cases and 155,000 deaths occur each year from gut infections and diarrhoea. A further 3.4m cases of invasive infection are estimated to have a much higher death rate - 681,000 annually - with the highest incidence among the 1.9m cases in Africa.
EI has been working jointly with the Quadram Institute and the University of Liverpool to research Salmonella and its impact on human health. The work has made use of EI’s genomics platforms, systems biology expertise and high-performance computing capabilities to perform a systems-level comparison of five Salmonella strains that cause problems in the human gut with five strains that infect organs outside the gut such as the liver and spleen, among others.
EI and QIB researchers have been looking at the genetic differences between these and less invasive strains.
Understanding the mechanisms that have allowed some Salmonella strains to invade other organs is the first stage in developing new drugs or approaches that can treat such infections.
The research to overcome the current limitations of the field has led to the creation of SalmoNet.org, an open access, dedicated resource for Salmonella data. This allows strain comparisons and enables detailed modelling, host-pathogen interaction studies, novel drug discovery approaches, and experimental testing to uncover thus-far hidden mechanisms, which could lead to developing new treatments for Salmonella infections.
Unlike using general antibiotics which may kill several other bacteria, and often could contribute to antibiotic resistance, this approach could support the development of smart drugs targeting strain-specific proteins in the infecting Salmonella.


Unlike using general antibiotics which may kill several other bacteria, and often could contribute to antibiotic resistance, this approach could support the development of smart drugs targeting strain-specific proteins in the infecting Salmonella.

A complementary project involving also EI and QIB has produced molecular maps of gut cells of different types to understand which become infected by Salmonella and how. Understanding how an invading pathogen like Salmonella works could identify host proteins targeted by these pathogens, and lead to the development of treatments to counteract the pathogen.
The EI and QI teams are applying the results of the research to probiotics with the aim of understanding which ‘good’ bacteria are beneficial to the gut and why. This research could support the development of new, more effective probiotics, or it could support health claims for existing probiotics, creating substantial market value for the companies involved.
If successful, EI’s impact globally could be £1.1bn over the next 25 years.

A rampant disease caused by the fungal pathogen Hymenoscyphus fraxineus, ash dieback has spelt doom for much of Europe’s native ash trees and, since 2012, has been wreaking havoc in the British countryside. It’s estimated that up to 95% of trees could succumb - figures already seen in some areas of Scandinavia.
EI was part of the rapid response to ash dieback in the UK, forming part of the Nornex consortium established in 2013 to research both the pathogen and ash trees through genome sequencing and population genetics approaches, hoping to find the genes that make the pathogen a success, as well as those that confer resistance (or tolerance) to the pathogen in some ash trees.
Well annotated genome sequences tell us a lot about how we might identify native ash trees that show some resistance to the disease, or can accelerate the process of breeding some much needed resistance into new trees to help transplant some further defence against the onslaught, though more recent studies point to an even greater threat.
Subsequent research on the ash dieback pathogen, led by Earlham Institute, found huge variation in the pathogen in its native range in the Far East of Asia but much less variation in Europe, suggesting that increased variation in the pathogen here could be even more devastating to ash populations and that on-going import protection and surveillance are very much worthwhile.

EI’s research found that the devastating invasion of ash dieback arose from just two individuals, meaning that the introductions of just one more spore could spell doom for those 5% of ash trees that remain. The reason that this is so potentially devastating is that there are many genes that can turn a fungus into a disease-causing pathogen, many more of which are present in the native Japanese range. This might add to the effectiveness and destructiveness of the current European pathogen if a spore was to arrive and introduce more of these genes.
There is strong public interest in protecting woodlands, particularly those with species so iconic to the national landscape and important for biodiversity as ash - a tree on which thousands of other species live and many require absolutely to survive. Brookdale Consulting estimate a total value of ash in Great Britain of £655m per year taking account of economic, social and environmental values.
Safeguarding this resource is valuable for the UK and in light of this, it’s estimated that EI’s contribution to understanding this pathogen threat will amount to £16m over the next 25 years.
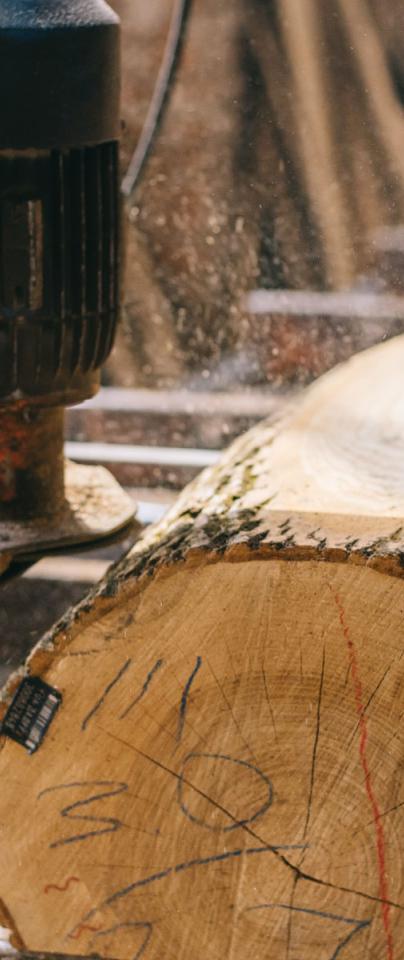


There is strong public interest in protecting woodlands, particularly those with species so iconic to the national landscape and important for biodiversity as ash - a tree on which thousands of other species live and many require absolutely to survive.

These projects are highlights of what we’ve done over the last decade, but we have recently taken on many more projects that have seen our global impact really push on.
One example is our leading role in the GCRF-funded GROW Colombia project that aims to preserve biodiversity and increase socioeconomic well being in the country in order to achieve a sustainable peace while establishing sustainable and lasting research infrastructures.
Another prominent project will be the Darwin Tree of Life work that we are undertaking alongside partners from the Sanger Institute, NHM and beyond, which, as part of a broader, international, Earth BioGenome Project aims to sequence all known life on Earth.
