Salmonella, who gives a faeces?
Salmonella is not just about bad eggs, it causes a devastating global disease to rival malaria. How is bioinformatics helping to tackle it?
Salmonella is a prevalent global pathogen, killing over 150 000 people each year (sometimes even up to one million). Researchers at the Earlham Institute are helping to understand this important bacteria through understanding its invasiveness, pathogenicity, genetics and evolution, especially with regard to the growing threat of multidrug resistance and alarmingly devastating new strains arising in sub-Saharan Africa.
Particularly of interest to EI scientists is the species Salmonella enterica, and more specifically still, Salmonella enterica serovar Typhimurium. This is just one of the 2,500 types of salmonella (which are all very pathogenic - we’ll come back to this later), but is remarkable for just how widespread it is, infecting a wide diversity of animal hosts throughout the world.
It’s not just a pain in our intestinal wall, it’s also prevalent in livestock and wild animals - hence the need to cook your chicken properly, or wash your salad (that may have been handled by someone who has handled some raw meat, or other contaminated product), lest you want to spend a shitty week clutching at your abdomen and chundering from either end through fits of fever and chills.
Though most afflictions due to Salmonella are self-limiting cases of gastroenteritis, it is a big problem in the developing world - causing up to 400 000 deaths in immunocompromised patients in sub-Saharan Africa, for example - where Salmonella variants have adapted ways to get into the bloodstream.
There is also the encroaching, and quite terrifying, issue of multidrug resistance, which, increasingly, will hamper efforts to tackle Salmonella - and other widespread bacterial pathogens - in the future.
So, how are researchers at the Earlham Institute, through important collaborations with the Quadram Institute and the University of Liverpool, helping to understand Salmonella through its diversity, pathogenicity, adaptation and evolution?
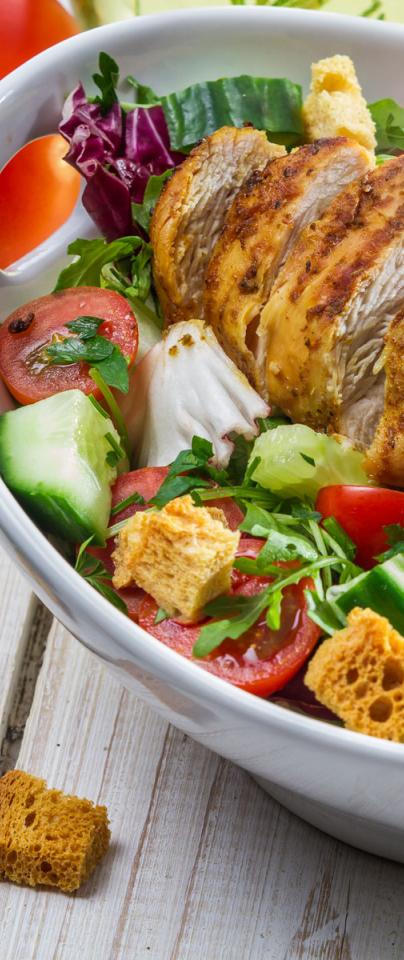


It’s not just a pain in our intestinal wall, it’s also prevalent in livestock and wild animals - hence the need to cook your chicken properly, or wash your salad, lest you want to spend a shitty week clutching at your abdomen and chundering from either end through fits of fever and chills.

Salmonella, a group of bacteria, is most genetically similar to E. coli, which might not come as a surprise, as E. coli is a species of bacteria found in guts worldwide, mostly living happily alongside the other microbes that make up a healthy gut microbiome.
E. coli can get nasty, however, as is well documented - with lethal food poisoning cases involving lettuce in recent years, among others - but mostly, E.coli strains don’t threaten our guts. That’s where salmonella differs.
All types of Salmonella are pathogenic (disease-causing) due to a number of “pathogenicity islands” (we’ll explain what these are in a minute) that were picked up early on down the evolutionary split away from its most common ancestor with E. coli.
These Salmonella pathogenicity islands were gained via horizontal gene transfer - transfer of genes not from “parent” to “offspring” - which, incidentally, is the primary mechanism for the spread of antibiotic resistance (which we’ll come back to later). Bacteria essentially have a tube which they use to transfer bits of DNA between one another through a process known as bacterial conjugation. (Bacteriophage viruses can also act as the vectors, which is for another blog post!)
Yes, bacteria have sex, too, and it’s weird.
Horizontal gene transfer doesn’t even have to be between the same species. Salmonella can pick up bits of DNA from E. coli or other bacteria (the concept of the “species” in bacteria is actually very loose and difficult to define, salmonella being a perfect example of this).
Those horizontally acquired “pathogenicity islands” are groups of genes (see our article, “what’s in a gene”) coding for “virulence factors” (i.e. proteins that help disease-causing bacteria infect something) which are not present in closely related species that are not pathogens. Salmonella enterica has picked up two defining pathogenicity islands (actually many more than two overall), making it a pretty successful gastrointestinal pathogen - although, it doesn’t limit itself to guts and is starting to wreak havoc by invading the bloodstream of species that it has become particularly well adapted to.
Of all of the varieties of Salmonella enterica, Typhimurium is the most interesting to look at because, although other serovars (posh word for varieties) are found in all sorts of animals, this one is seemingly among the top two found in all animals that we’ve cared to study thus far. Some serovars, such as Salmonella enterica serovar Gallinarum, affect only birds, while others, such as the serovar Dublin, only affect cattle.
S. Typhimurium affects pretty much every unfortunate gut it comes into contact with.
And what it does to those poor guts!

Yes, bacteria have sex, too, and it’s weird.

Tricky little bacteria, in the right conditions - usually the lower intestine - Salmonella becomes an utter menace to the guts of animals.
It requires the right sort of environmental triggers, which might be the types of small chain fatty acids, or the relative oxygen levels, for example, but once triggered Salmonella puts into effect its arsenal of weapons that confuse our body’s own mechanisms that normally help keep us healthy.
For a start, intestine cells are not phagocytic, i.e. they’re not like the white blood cells that patrol our bodies and gobble up harmful-looking bacteria. The first thing that Salmonella does, once it has attached to the gut epithelial cells that make up the inner lining of the small intestine, is use a bacterial syringe-like apparatus (called a type 3 secretion system) to inject molecules into gut cells, which trick them into becoming phagocytic so Salmonella can get inside those cells.
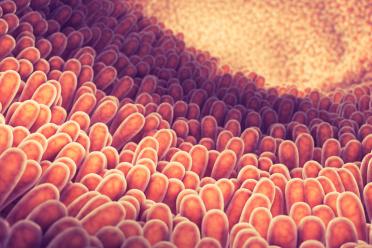
Imagine your intestine, with its inner wall made of lots of little folds (villi), which normally help to digest food. Salmonella manages to convince cells in those folds to grow and form a protective envelope around the invading bacteria, which essentially means that it becomes one with an epithelial cell in the intestinal cell wall through remodelling the cell cytoskeleton (the proteins that give cells their various shapes and forms).
This protective envelope essentially means that the Salmonella cell is now protected inside the gut wall in a sort-of vacuole (a storage compartment within a cell), where the bacteria then injects more proteins (using another molecular syringe). This similarly confuses the body’s normal processes such that, in a scene reminiscent of some sort of alien-horror movie, this protective vacuole, now hidden from the body’s defence mechanisms, extends outwards with projections that expand the space within the cell for the bacteria to multiply.
This all leads to the inflammatory response that causes up to a week of diarrhoea, nausea and vomiting that are associated with Salmonella infection.
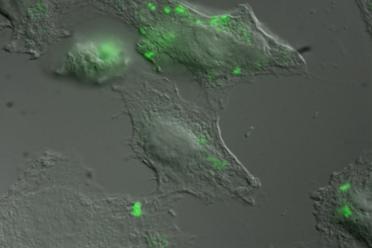
At Earlham Institute, the Korcsmáros Group is helping to understand more about how Salmonella infects and controls our gut cells using models such as organoids (very cool systems based on stem cells that can be turned into tiny models of a human gut), and techniques such as fluorescent labelling (see the images provided by Isabelle Hautefort). Combined with computational analysis and development of tools, such as SalmoNet, this allows our scientists to analyse the multiple, complex interactions between Salmonella and its animal hosts (like us).
Isabelle Hautefort and Amanda Demeter from the Korcsmáros Group are working on a very interesting project alongside Iain Macaulay’s single cell group at EI. Iain’s group have some pretty amazing kit that can separate single cells, and then sequence the DNA of these individual cells, so that we can compare single gut cells infected with Salmonella and ones that are not.
In this way, we can better understand what’s going on in the infected cells by seeing what genetic changes have taken place. Better still, we can take a look at what happens in different types of gut cells, including Paneth cells, which are a very rare type of cell in the gut involved in the innate fighting response to microbes yet can still be targeted and infected by nasty bugs like Salmonella.
We can predict and validate which Salmonella proteins, produced during infection, are likely to bind to which host proteins, affecting entire host cellular processes and allowing the pathogen to hijack its host’s cell machinery for its own benefit. One important host cellular target of Salmonella among other pathogens is autophagy, an important cellular clearance process essential to recycle damaged organelles and misfolded-proteins, as well as clear intracellular pathogens like Salmonella.
A quite scary development in Salmonella, however, is that some varieties of Typhimurium (that’s right, there are varieties within varieties), which are generally adapted to a wide range of hosts, have become more host-adapted (specific to one animal), and have developed the tendency to spread from the gut into the bloodstream, causing deadly, systemic infections, particularly in sub-Saharan Africa (albeit in immunocompromised patients).
More terrifyingly still, these strains (invasive non-typhoidal salmonella - iNTS) are also showing high resistance to multiple antibiotics, which renders them very difficult to treat (more on that later).

More terrifyingly still, these strains are also showing high resistance to multiple antibiotics, which renders them very difficult to treat.

In the UK, we have been sequencing all of the Salmonella strains that the NHS come across for a number of years, and we have a very good picture of the different strains and genotypes of Salmonella that affect UK patients (over 200 000 European Salmonella genome sequences are stored in the University of Warwick’s Enterobase).
This information has not been so available for people in the developing world, which is all the more worrying when the mortality rates for Salmonella in sub-Saharan Africa, especially, have been staggeringly high. Proper treatment of infections is greatly influenced by a better understanding of the specific genotype, or strain, of Salmonella involved. Hence, Neil Hall - director of EI - along with Jay Hinton of the University of Liverpool, has been leading the UKRI-funded 10,000 Salmonella Genomes project.
The project is funded to sequence and analyse 10,000 Salmonella genomes from various countries in Africa and Latin America, including Colombia, Mexico, Malawi and Uganda. The information gleaned will hopefully provide insights into how the bacteria spreads, the emergence of drug resistance, and the virulence factors that render the bacteria such a global threat to human lives. Particularly of interest, two distinct lineages of S. Typhimurium ST313, which cause the highly fatal bloodstream infection we’ve already brushed on.
The sequencing of all of these Salmonella genomes has already taken place at the Earlham Institute, using Darren Heavens’ innovative LITE pipeline - a low cost, low input, automated method for generating genome sequences quickly.
Chris Schudoma of the Swarbreck Group at EI then helps to assemble the bacterial genomes, while Ross Low of the Neil Hall Group then puts the cherry on the top - finding the interesting information within all of those genomes among the genes and other elements that make Salmonella such a menace.

The sequencing of all of these Salmonella genomes has already taken place at the Earlham Institute, using Darren Heavens’ innovative LITE pipeline - a low cost, low input, automated method for generating genome sequences quickly.

The collaboration has recently delivered a paper, led by Yan Li of the University of Liverpool who performed the analysis of several of the genomes sequenced at EI, which focuses on non-typhoidal Salmonella (Typhimurium) infections in Colombia (responsible for 32.5% of infections in the country), which delivered some interesting and important results, especially considering the lack of information on bloodstream-associated Salmonella previously in Colombia. Among the findings were the first isolates of ST313 to be reported in Colombia.
Another very interesting result was that one of the seven clusters of Salmonella identified from the analysis, which tracked 209 Salmonella genomes gathered between 1999 and 2017, was related to a European multidrug-resistant strain.
Among the genes identified in the study, the team was also able to link resistance to a drug - nalidixic acid - to the qnrB19 gene.
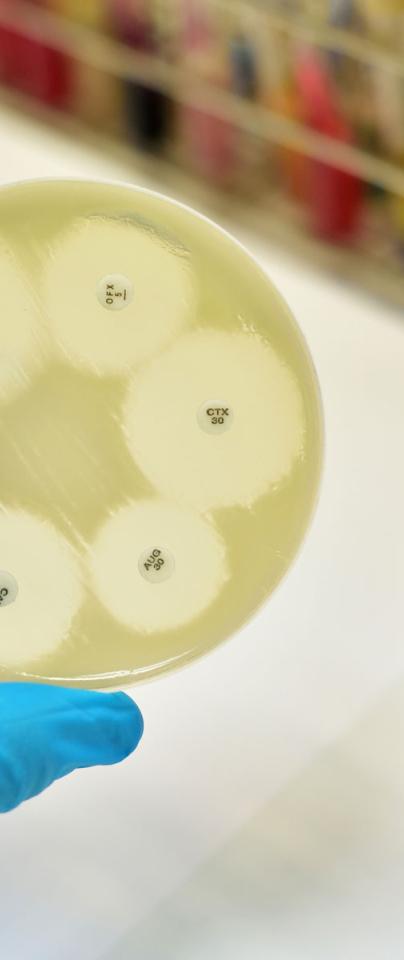

Bacteria are sharing bits of each other’s DNA all the time, and it’s a problem when it comes to treating bacterial infections, especially when we haven’t discovered a single new class of antibiotics for decades.

It is the multidrug resistance, as well as the global spread of such invasive strains of Salmonella, which is of great cause for concern, but also of great use for scientists wanting to understand how bacteria are evolving to get around our medical interventions.
Another EI (and QIB) researcher, Matt Bawn, is particularly interested in antibiotic resistance - particularly how low levels of antibiotics present in the environment might be contributing to the crisis.
We already mentioned the pathogenicity islands that Salmonella inherited after it split from its most recent common ancestor, E. coli, which make it pathogenic. Similar bits of DNA, swapped between bacteria, are also at play when it comes to antibiotics.
Bacteria are sharing bits of each other’s DNA all the time, and it’s a problem when it comes to treating bacterial infections, especially when we haven’t discovered a single new class of antibiotics for decades. The fight against antibiotic resistance is one of the biggest fights we’ll have over the coming years: it’s not so many years since TB, the consumption, caused by the bacteria Mycobacterium tuberculosis, was culling people in their millions.
The routes by which bacteria obtain resistance to drugs, as well as how they obtain the factors required to become pathogenic, are very interesting to study, as they don’t follow the usual routes that we might normally associate with evolution. Instead of slow, incremental changes to DNA sequences, bacteria tend to adopt (or lose) whole stretches of DNA sequence at distinct points in their evolutionary path. This challenging task is being aided by a BBSRC-TRDF grant recently awarded to EI and partners, with Iain Macaulay and Johana Hernandez of EI’s single cell sequencing team looking at the effects of growing bacteria in low level antibiotics and examining the effects.
Salmonella is a great model to study antibiotics resistance due to its global prevalence, as well as the fact that it has been so sufficiently sequenced, not just in Europe but around the world, thanks to projects such as 10,000 Salmonella Genomes.
An interesting example of how this can arise is found in the multidrug resistant pandemic S. Typhimurium strain which is responsible for around two thirds of the current global Salmonella pandemic. This strain contains several genetic regions of interest that confer resistance to a number of antibiotics, but one in particular contains genes thought to allow Salmonella to tolerate copper.

Salmonella is a great model to study antibiotics resistance due to its global prevalence, as well as the fact that it has been so sufficiently sequenced, not just in Europe but around the world, thanks to projects such as 10,000 Salmonella Genomes.

Why is this interesting?
This region has been also linked with the success of another Salmonella strain found in pig herds in which copper is used as a growth promoter due to its antimicrobial properties. Thus, the bacteria present in the pigs have become resistant to the use of copper as an antimicrobial.
This is tremendously important knowledge and a case in point: our low level use of antibiotics, and other growth enhancers, in livestock, fish farming and elsewhere, are most likely a big cause of multidrug resistance in bacteria - and these bacteria are sharing their drug resistant regions with each other.
Matt has an analogy that he likes to use, which explains well the phenomenon.
If we imagine that the Earth is a tiny sphere relative to a huge giant which stands above it with a huge glass of water the size of Mars, then the glass of water will drown all the humans. However, if the giant feeds in low levels of water, then the people will use it and probably thrive.
As with imaginary globes of drowning people, as with real bacteria all over the planet, which are adapting rather well to the compounds that are supposed to kill them.
It’s not just drugs that bacteria are developing resistance to, but even hospital cleaning products, for example, which also exert a ‘selection pressure’. Anything that the world throws at them (in the name of killing them), evolution finds away for them to get around.
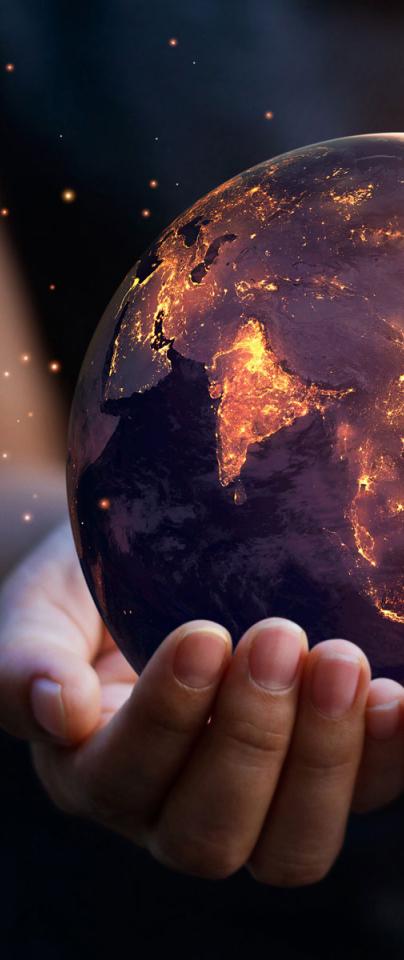


If we imagine that the Earth is a tiny sphere relative to a huge giant which stands above it with a huge glass of water the size of Mars, then the glass of water will drown all the humans. However, if the giant feeds in low levels of water, then the people will use it and probably thrive.

Salmonella, then, is a tricky little group of bacteria - and one that is constantly adapting, changing, and throwing out new ways to become even more of a menace to society.
Hopefully, the research being undertaken by EI scientists and their collaborators, spanning all of the projects we’ve explored in this article, will help to better understand the spread, virulence and evolution of Salmonella.
This information could help to serve global efforts to kerb the devastation that it continues to serve in our guts, and might also inform our research into other diseases along the way.
Earlham Institute receives funding from the Biotechnology and Biological Sciences Research Council (BBSRC) to provide National Capabilities and to deliver Strategic Programmes.

A dedicated, high-throughput genomics facility is out of the reach of most organisations due to the large capital investment and the requirement for skilled, multidisciplinary staff. Small-scale investment in genomics platforms in an institute or research group cannot deliver UK research ambition for high-throughput, massive-scale studies on single-cell systems or populations.
The National Capability in Genomics and Single Cell analysis delivers a cost-effective facility to assay genomic diversity, while extending this with a critical national research capability in high-throughput, multiomic, single-cell analysis and addressing a very current need.
This National Capability will continue to provide researchers with access to new technologies, bespoke bioinformatics and high-throughput lab pipelines with a focus on crop, microbial and non-model animal genomics, delivering a strong scientific impact and strategic importance in the area of agri-science, in particular.
The complementary genomics platforms and expertise of the EI Science Division supports this National Capability to address complex genomics questions in agriculture and food security. Closer links to the Norfolk and Norwich University Hospital and Quadram Institute will transition this expertise to bioscience for health, particularly in microbiome studies of health and nutrition.

Understanding how living systems work, across interactions at the micro and macro level, is crucial for a deeper understanding of the biology of organisms. We will deliver impact across food security and sustainability through the development of predictive and testable models in economically important organisms. Industries our work will support include aquaculture, plant biotechnology and pharma. Our work will be made available to the wider research community through publications and open data resources to maximise knowledge exchange opportunities.