Orchestrating life: miRNA
How do small pieces of RNA help determine how genes are expressed?
Genes are just a part of the genetic score, units of information that give us (sometimes) the proteins that eventually help make, carry, transport or break other important molecules. Sometimes genes are “turned on”, and sometimes they are “turned off.” Yet genes are not binary. The orchestra of life is a finely tuned system.
The bright orange root of a carrot looks a lot different to its green leaves, yet the cells which make them both up carry the same DNA. It’s the same for our blood, nerves, eyes, livers, spleens, muscles, hearts, brains and bowels - in any one individual, 100% the same genes.
The human body manages to produce at least 200 different types of cell using the same instructions, so there must be something else going on.
That’s where things such as microRNA (miRNA) come in - and scientists at EI are using computer-based bioinformatics methods that are helping us to understand their role in the evolution of different characteristics in mammals.
The human body is made up of 200 types of cell, which all use the same instructions to differentiate into all of their diverse forms. MiRNA is one of the genetic mechanisms that ensures DNA blueprints aren’t all translated the same way.
You can perhaps think of life as an orchestra, which is composed of dozens of different instruments - violins, cellos, pianos, drums, oboes, trumpets, bassoons and more. If all of these were to play all at once, at full volume, the audience wouldn’t stick around very long, and they certainly wouldn’t give us what Tchaikovsky meant us to hear.
The instruments need to be tuned, played along to different melodies. Sometimes they are not being played at all, other times they are all playing at the same time but with different pitch and amplitude, louder or more softly, staccato or legato.
To guide the orchestra through this myriad of potentialities is the conductor, who guides each musical section along to help produce the melodies that melt our ears. Cells need a conductor, too, and in this case it’s miRNAs.
We often hear about genes being turned on or turned off, or even edited out. Yet, this only tells a small part of what is going on. Gene expression can be better understood as more akin to a dimmer rather than a simple on/off switch; the lighting (or expression in this case) can be finely adjusted.

The reason that this works is due to the nature of how DNA is read by cells so that they can make the appropriate proteins. DNA is a double stranded molecule that has a single stranded relative, RNA. The first step in reading DNA is called transcription, when a DNA sequence is copied to form a stretch of RNA called messenger RNA (mRNA).
Lots of other things happen at this point, such as alternative splicing, which is where one gene can make lots of different proteins depending on which parts you stick together. However, that’s a topic for another post. The important thing is that DNA gets converted to RNA.
At this point, the RNA is then taken to one of the millions of ribosomes where the instructions are then read and translated into a protein code, in the form of amino acids which are delivered to the ribosome and stuck together like LEGO to make proteins.
Where miRNAs come in is in between the transcription and the translation (or sometimes at the stage of translation itself).
MiRNAs are tiny, just 22 chemical letters long. These 22 letters, in the case of a large number of messenger RNAs, just so happen to be a perfect complement to specific sequences so that, just like with DNA, they can bind to a stretch of mRNA.
When the miRNA attaches to the mRNA in this way, the cell is then instructed to stop protein construction and sometimes even destroy the mRNA. In animals what happens is mostly translational repression, while in plants miRNA regulation almost always leads to target cleavage.
Obviously, this effect is not 100% effective, hence the “dimmer” effect. Some miRNAs are themselves more highly expressed in certain cells, so that, in the brain, for example, the same protein might be made but in different amounts compared to the same protein in the heart.
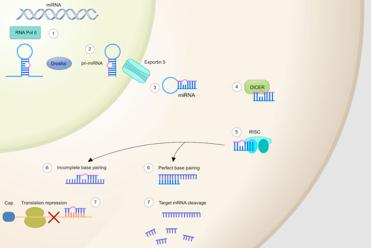
MiRNA is single stranded, but exists in a “loop” whereby some of the matching chemicals can still bind to each other - it is copied from DNA in the same way that genes are copied into RNA, in the nucleus of the cell (1). Eventually the miRNA is transported out of the nucleus and into the cytoplasm of a cell as pri-miRNA (2) and is then converted to miRNA by an enzyme called DICER (3,4), which helps to activate the RNA-induced silencing complex (RISC) (5), which then, using the miRNA as a guide, targets the correct mRNA (6) and destroys it or represses translation (7).
What better way to help explain the wonders of miRNA regulation of gene expression than by looking into the latest research coming out of the (former) Di Palma group at Earlham Institute, where Luca Penso Dolfin has been investigating the role of miRNAs in mammals.
His latest work has been to look at the role of miRNAs in the different tissues of dogs, pigs, cows, horses and rabbits - namely the brain, testis, kidney and heart. The research, “The evolutionary dynamics of miRNAs in domestic mammals”, was published in Scientific Reports this year.
Why would we want to study such animals?
Well, for a start, all of these animals are important for economic and biomedical reasons. The economic value of cows and pigs is obvious. Biomedically, knowing more about how miRNAs affect development in pigs, for example, which are anatomically and physiologically very similar to us, might help us understand more about ourselves.
Aside from these examples, in general we can learn much more about ourselves by looking at what happens in all other genomes than we can by studying ours alone. Identifying the miRNAs that are conserved, and those that are different, between different lineages, offers a fascinating insight into the evolution of specific lineages - especially those that are so similar, genetically speaking, as we are to our mammalian cousins.
It’s often stated that we share 98% of our protein-coding genome with chimps - and the similarities on the anatomical scale are incredibly obvious. We still share 85%(ish) with mice, 80%(ish) with a cow, etc., therefore it’s interesting to study the small differences that make us each unique.
Anyone who’s seen a picture of a human embryo compared to that even of a dolphin will know that, at the very early stages, there’s little to distinguish us from even our most distant relatives.
Developmentally speaking, miRNAs have an incredibly important role to play.
In humans alone there is at least one miRNA for every ten genes, with miRNAs thought to regulate at least a third of human genes. Interestingly, these miRNAs can exist in the genome as either non-protein coding genes in their own right, or as part of a non-coding region within a protein-coding gene.
This, for one, shows how it’s very difficult to regard any DNA sequences as “junk”, and on the other makes a lot of sense. It wouldn’t perhaps work very well for a protein-coding, transcribed gene region (an exon) to include a miRNA, as that would mean the mRNA would be cut up and lost.
Indeed, the recent EI study highlighted the importance of “intronic” regions (the non-protein coding sections of genes) in the emergence of novel miRNAs, with 366 sets of sequences found. Duplication events and repetitive elements contributed another 135 and 37 sets of sequences, respectively.
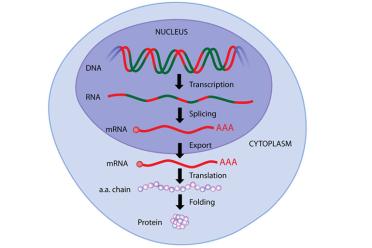
A eukaryote (human, animal, plant, fungus, alga etc.) gene is made up of protein coding “exons” (red) and non-coding “introns” (green). The introns are chopped out before a protein is made. Many (if not most) miRNAs, as well as other non-coding elements such as long non-coding RNAs, are found in these regions, which you can read about in another of our articles.
Looking across the five mammal species, the EI team identified 2088 miRNA sequences, 412 of which were novel, or “new”, all based on computational predictions followed by manual curation through shrewd bioinformatics analysis.
A whopping 20% of the novel miRNA sequence groups were found only in the brain, found to most likely regulate neural processes, among other things. The next most likely location was the testis, which might not come as a surprise when talking about evolution.
Considering the myriad of shapes and forms of dogs, especially, as well as the other domestic animals that we have selectively bred in just about 10 thousand years, it’s clear that there is a relatively quick path towards evolutionary changes at the phenotype level - one that is likely caused in no small part by miRNAs.


Dogs come in all shapes and sizes, and have been selectively bred to appear this way in just ten thousand years or so. Perhaps it is miRNAs that are behind many of their physical and behavioural adaptations to life as man’s best friend.

Out of all of the animals analysed, it was the dog which showed the highest number of novel miRNA genes, with 166 out of 547 detected. Even taking into account that the dog genome is better sequenced and more brain samples were taken, the dog came out on top.
Indeed, the dogs and cows, in particular, showed a high emergence of species specific “seed sequences”. Seed sequences are the bits of miRNA that match exactly the sequence of the mRNA that they target for downregulation.
Interestingly, these species specific seed sequences have target genes that can be found mostly to have a neurological, behavioural or immune related function, which has fascinating repercussions for miRNAs in the domestication process, as all of these processes are positively selected in domestication.

All in all, miRNAs represent “a relatively simple source of innovation”.
Most miRNAs are indeed conserved over long time periods, but these match very well conserved target sites. However, there is a high turnover rate of miRNA seed sequences, while these sequences are so short that there is a high chance of targeting different mRNAs and therefore offering a range of regulatory changes in relatively short time frames.
Though a concrete role in shaping the adaptation of specific species of animals is still to be unearthed, the specificity of miRNAs identified in this study points towards a role for miRNAs in shaping the evolutionary path of domesticated animals. It will be interesting in the future to compare domestic species with their wild counterparts to further probe the effect of selective breeding, and the overall effect of miRNAs in the evolution of novel regulatory pathways and phenotypes.