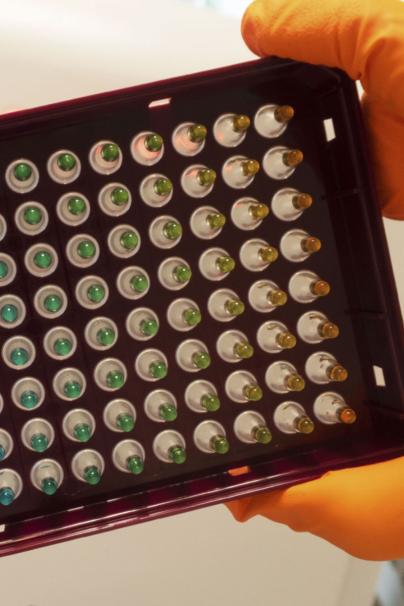
Single-cell genomics can provide very detailed information and enable us to look at tissues at an unprecedented resolution. We are able to define which types of cells compose a tissue and how they work together to develop and conserve the tissue during an organism’s life.
Moreover, we can describe what changes happen when a disease initiates and progresses - and in which most sub-groups of cells within the tissue, these changes occur - whereas this information remains hidden when performing a bulk study. This kind of analysis can be applied to every tissue and organism; to study meiosis; how a tissue develop during embryogenesis; or which group of cells in a tumour develop resistance.
My project at EI aims to develop novel single-cell sequencing approaches that provide more information about how transcription, the process to translate DNA into RNA, is regulated and apply the approach to study blood stem cells. In particular, to define if blood stem cells are heterogeneous and primed to differentiate into a specific type of blood cells, and how the stem cell makes the decision to start differentiating. As blood stem cells seem to lose their ability to properly replenish blood tissue when the organism ages, we are applying this approach I developed to studies where cell changes may cause this loss of function.
I have been fascinated by biology, in particular genetics, since I first studied it at high school. I decided to study biology at University; during that time and throughout my PhD I became really passionate about molecular biology and research. I got into single-cell genomics gradually, moving into genomics first during my postdoc, when I had the opportunity to learn about all the different technologies and their potential to study disease. I was really fascinated by the idea of looking into a tissue cell-by-cell and finding out how they maintain the tissue, and what happens when disease occurs.
In my science career, one of the biggest challenges is to learn to deal with failure. Science requires time and dedication and sometimes you may not get the results or the output you expected. It can also be quite difficult to balance a personal life with non-permanent jobs and the need to move to a different lab and city, or sometimes country. It is really rewarding, on the other hand, contributing to advancing scientific knowledge and hopefully making an impact. My ideal research project would be one in which I could develop a novel technology and use it to answer a crucial biological question.
I feel gender equality is becoming better, however, I think there is still a gap in more senior positions. In the future, I would like to keep working in research to gain more knowledge and scientific independence - the best thing you can do is to be really passionate about your science and don’t be afraid to ask your peers for help and suggestions.