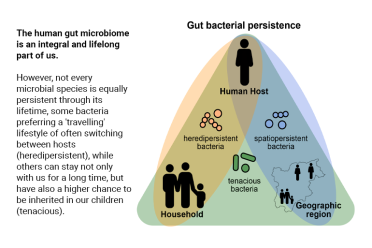
Researchers in the UK and Germany, alongside other international collaborators, have investigated the evolution of bacteria in the human gut microbiome - asking how these microbes persist throughout their lifetimes - taking into account internal and external influencing factors.
The results of the study will help inform tailored probiotics, live bacteria found in particular foods or supplements, as well as dietary or medical interventions to treat gut disease and maintain a healthy gut microbiome.
Keeping a stable, healthy gut microbial population is mutually beneficial to us and the bacteria. In exchange for nutrition and a comfortable habitat, the microbe community returns the favour by providing us with health benefits, which we are now starting to understand.
Lead author and Group Leader Dr Falk Hildebrand from the Quadram Institute and Earlham Institute, explains: “We know that certain microbes colonise us at birth and some can live with us for decades. Yet, although studies have looked at individual microbe species, the mechanisms and scale of persistence in the microbiome as a whole haven’t been explored.”
To examine this, a team of scientists at the Earlham Institute and Quadram Institute on the Norwich Research Park, along with the European Molecular Biology Laboratory (EMBL) in Germany, used metagenomics to analyse the evolutionary strategies and persistence of different bacteria in the human gut microbiome.
Metagenomics is the study of all of the genes from many different organisms in a population. In terms of the human gut microbiome, this process not only provides detailed information about the bacteria strains present but also indicates the enhancing capabilities of those different strains, based on their genetics, to keep the gut in good working order.
From analysing stool samples, the team re-examined metagenomes from over 2,000 adult and infant samples, including several from the same families and found three major dispersal strategies underlying human gut bacterial persistence. The data came from previously published studies looking at microbiome changes over time, with each individual providing on average 2-3 samples several months apart.
Last author and Director EMBL Heidelberg (Scientific Activities) Prof Peer Bork, said: "By looking into time series from individuals and family members and overlaying this with geographic information, ranging from household via city to country, we identified groups of bacterial strains that show different dispersal strategies. This presented very different persistence patterns in the host, regional spreading and the geographical distributions of hundreds of bacterial species."
The data was built into a diverse dataset of 5,278 metagenomes, which were probed to analyse patterns of persistence in the different types of bacteria and how these were influenced by the common factors: age, family members, geographic region, and antibiotic usage.
“Our analysis shows that most strains of bacteria present in the microbiome are very persistent - with the chances of a strain persisting for at least a year being over 90%,” said Dr Hildebrand.
“Some microbe species did show consistent differences being either highly persistent taxonomic groups, or being low-persistent, relying more on exchanges between family members. In babies, however, the average persistence of bacterial strains dropped to 80%. This isn’t unexpected; we know that especially in newborn babies there is an ongoing exchange of gut microbes.”
Prof Bork, added: “What the study shows is that the intrinsic persistence levels of bacteria seen in adults are also reflected in children, and gradually we start to acquire those persistent bacteria up to about ten-years-old at which point the microbiome reaches a steady state. “
“Antibiotics had different effects of different types of bacteria, with the overall effect depending on how resilient different bacteria are, their intrinsic persistence, and to what extent they were replaceable within the microbiome.”
To delve deeper into what drives persistence, the researchers compared microbiome communities beyond an individual level, but also across families, countries and regions. This allowed them to group bacteria based on their persistence characteristics and, through genomic analysis, look for clues to the evolution of these groups’ strategies in dispersing among new human hosts.