Genome sequencing comes in many forms nowadays. Though it’s more accessible than ever, achieving high quality still requires the right combination of talent, tools, and experience.
At the Earlham Institute, our Genomics Pipelines team and Haerty Group have applied all their expertise to assemble the best tilapia reference genome yet, which is helping WorldFish to breed more resilient and higher producing fish.
“It's about finding a solution with our pipelines and services, applying our technology to a scientific question,” says Dr Chris Watkins. He leads the project management team in Genomics Pipelines - the first port of call for any genome sequencing project.
“We make sure to understand what our customers and collaborators want, and apply the right solution in the lab.”
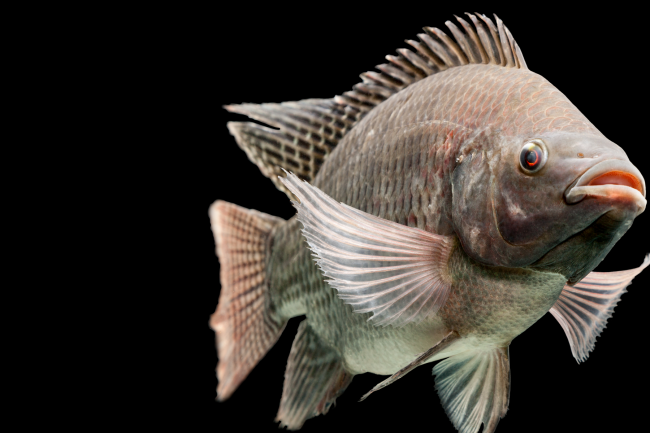
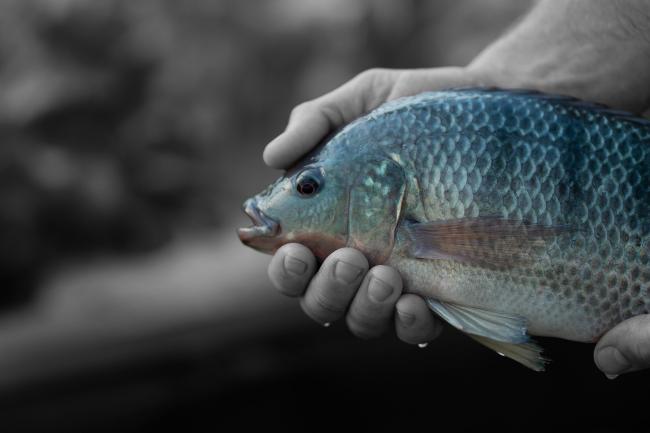
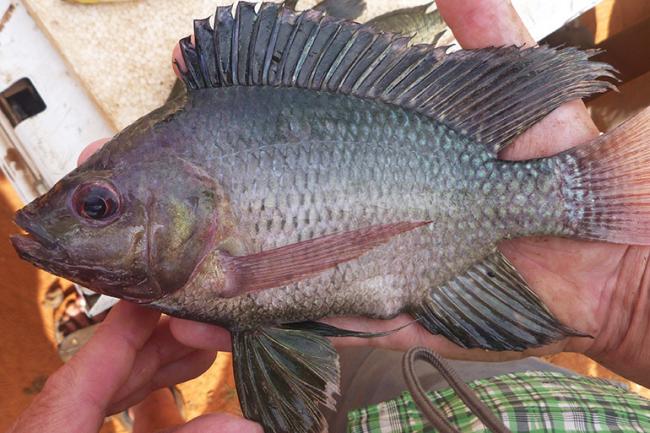
In the case of tilapia, researchers at WorldFish and the Earlham Institute’s Haerty Group wanted a high quality reference genome for GIFT (Genetically Improved Farmed Tilapia) - an elite variety of one of the most widely-farmed freshwater fish on Earth.
“You've got the full gamut,” says Watkins. “DNA and RNA extractions, Illumina PCR-free, 10x linked reads, PacBio HiFi, and RNA-seq. This was an opportunity for us to test and develop new protocols as well, so it’s been a nice project to work on.”
Here, we speak to the people doing each part of the process - generating high-quality genome sequence data to help the bioinformaticians put together the best possible genome assembly.
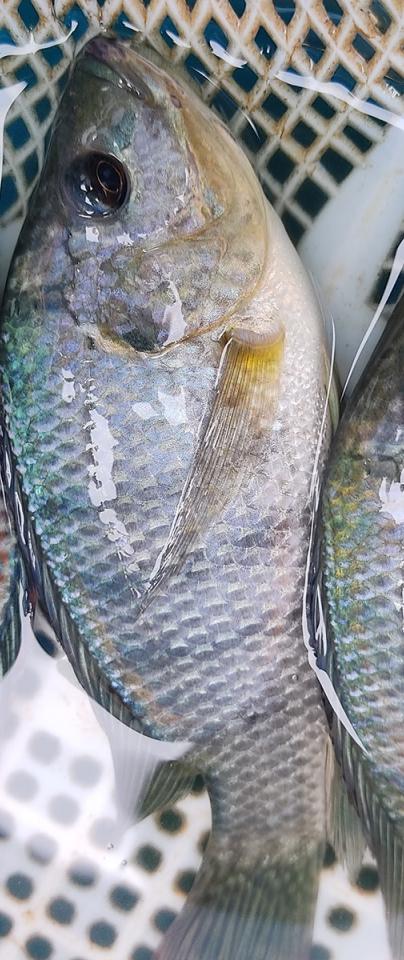


Because of the technologies applied, we should be able to further improve this assembly to chromosome length scaffolds and provide an assembly of even better quality than the current reference.

Dr Wilfried Haerty, Earlham Institute
DNA and RNA extraction
“The most important thing is to get good quality DNA to begin with,” says Genomics Pipelines Team Leader Leah Catchpole. “That’s where Alex comes in. He’s been fundamental in improving the DNA that we have coming into our genome assembly pipeline.”
For the sequencing of the GIFT tilapia genome, Senior Research Assistant Alex Durrant applied his expertise to get the best DNA he possibly could. However, finding the best tissue to extract DNA from was far from simple.
“Liver seems to work well with mice, but it didn’t really work with tilapia. We also tried other organs, including heart and brain, but the extractions proved difficult and results weren’t always consistent,” Durrant tells us.
“Then we tried gonads, which didn’t look great to start with because there was so much DNA bound to the extraction disc it looked like a jellyfish. We tried it again with a lot less material and it worked really well, so it’s now the go-to tissue for all our tilapia projects.”
The project called for long-read DNA, which is why Durrant used a relatively new extraction method developed by Circulomics that allows for high molecular weight DNA to be obtained.
“Circulomics wanted to overcome the shearing of DNA that can happen with other methods by using just one magnetic disk, which is a flat circle coated in silica,” Durrant explains. “It's very gentle. And what’s nice is that you can modify the method fairly easily, such as reducing the input as we did for the tilapia DNA extractions.”

The most important thing is to get good quality DNA to begin with, that’s where Alex comes in. He’s been fundamental in improving the DNA that we have coming into the pipeline.

Leah Catchpole, Team Leader, Genomics Pipelines

Quality Control
“Once Alex has processed the DNA, and he is happy with it, it comes into Quality Control ,” says Catchpole. “We carry out extensive QC to make sure we can give the researchers we work with - in this case the Haerty Group at Earlham - a full set of recommendations for how to proceed down the appropriate sequencing pipeline.”
For sequencing, Durrant and Catchpole explain that there are multiple factors to consider, and various pieces of kit that can help ensure you’ve got the best material possible.
“The first stage is the Nanodrop,” says Catchpole. “That tells us if there are any contaminants in the sample, such as phenol, degraded RNA, or various carbohydrates.”
“Based on that,” adds Durrant, “it’s possible to go back to the extraction and understand what you need to do, such as more clean-up, or taking more care during a certain pelleting stage, for example.
“We then move on to Qubit quantification, which is a fluorescence based assay,” Durrant continues. “We use a dye that intercalates with the DNA, which gives us a more accurate assessment of how much DNA or RNA we have.
“The final step is running the samples on the Femto Pulse. It separates DNA, like with gel electrophoresis, except it’s done through microfluidics rather than having to deal with a big slab of gel.
“With that data, we can visualise DNA to a high resolution, which tells us what percentage is above, say, 40 kbp or 50 kbp.”

It's about finding a solution with our pipelines and services, applying our technology to a scientific question

Dr Chris Watkins, Project Management Team Leader, Genomics Pipelines

Library prep
Armed with lots of high-quality DNA, it was time for the experts in library preparation to work their magic and prepare the material to be sequenced.
“We've thrown various library methods at tilapia to get different sorts of information,” says Senior Research Assistant Fiona Fraser. “From Tom and Vanda who run the sequencing through to project management, it really has been a whole team effort.”
A mixture of sequencing platforms was used - from the long-read PacBio and 10X Chromium to the shorter-read PCR-free Illumina Novaseq.
PacBio Sequel II Hi-Fi
“A Hi-Fi library is where we make a really tight, precise library around about 15 to 20 kbp,” says Naomi Irish, a Research Assistant in Genomics Pipelines who has vast expertise in using the Pacific Biosciences platforms.
“We want to start with something that's really intact, as large as possible so that we are confident that the DNA is unlikely to be damaged,” explains Irish. “Then we take that high-molecular-weight DNA and shear it to the right size.”
From this point, the sheared DNA goes through various steps, including DNA strand repair and the addition of a barcoded adapter, as well as an enzymatic step that removes any incomplete library fragments that don’t have adapters on both ends. A piece of kit called the SageELF is then used to fractionate the library so that fragments of the perfect size are selected.
“It was one of the first Hi-Fi libraries we made, other than control samples that we ran to validate the method,” Irish continues. “Because Alex’s extraction was so good, we were also able to make a continuous long-read library with the DNA, which was really useful in the final assembly.”

Illumina Novaseq PCR-free
“It's a combination of different library preps that will give us a really good assembly,” says Catchpole. “PCR-free will give us a more representative look at the genome, without the bias of PCR.
“The quality of DNA for PCR-free libraries is important but, compared with Pacific Biosciences, size is not so [important] because the first thing we do with PCR-free libraries is shear to 1 kbp in length.”
Preparing a good library for sequencing, even with good DNA going in, requires bags of experience and expertise due to the multiple considerations when preparing samples.
“It's a real balancing act,” says Catchpole. “I used to do an overnight ligation, which would give the barcoded adapters more time to ligate to the ends of the DNA, to increase the efficiency. If we’re lucky, we’ll get 50% of fragments with adapters on both ends.
“The issue we have is, the longer you leave the reaction, the more frequently we get what we call ‘adapter dimers’ where we adapters stick together. The sequencers love these because they’re so small and so easy to sequence. So we no longer do an overnight ligation.
“We take a hit on the yield of the library rather than giving the sequencer something it loves to sequence but is of no use to us in assembling the genome.”
10X
The 10X Genomics libraries, a technology that has since been retired, were prepared by Fiona Fraser and added an extra dimension to the final assembly.
“10X gives us sequence information over long distances - maybe 100 kbp or longer fragments - and we can build up a detailed picture about how genes are organised along the genome,” Fraser explains.
“It’s quite a complex protocol,” says Catchpole, who adds that the 10X Chromium is like having a blend of long-read and short-read sequencing. “We start with high-molecular-weight DNA but sequence it on the Illumina using linked reads with barcodes that allow us to reconstruct DNA fragments bioinformatically.
“It’s a great way of having long-read information using a short-read sequencing technology such as the NovaSeq.”

It's a combination of different library preps that will give you a really good assembly

Leah Catchpole, Team Leader, Genomics Pipelines
Running the sequencers
Once the libraries have been prepared, it’s over to research assistants Tom Barker and Vanda Knitlhoffer on the sequencing side. Here again, experience and expertise are invaluable in ensuring that the best possible DNA sequence is generated.
“When we get the samples, we’ll qPCR them to make sure we’ve got an accurate quantification of the molecules with both adapters required for clustering,” says Barker. “ Illumina sequencers are generally robust. The main thing to consider is that certain libraries will cluster at different efficiencies.
“Making the most of the Pacific Biosciences sequencers isa bit of an art as it’s not possible to check whether we have the correct SMRTbells on either end of the fragments and contaminants are more of an issue, so it requires a bit more finesse at the beginning. The company has released a new loading method recently that should make the loading of the samples more consistent between cells, which could be very useful.”
“The Sequel II can generate very long sequences”, says Irish. “Although the structure of the library is linear, it's also topologically circular. And because the polymerase, once bound to the library, is positioned on the bottom of a well, it can incorporate bases while the library passes through in a circular repetitive fashion.
“With the right conditions, the polymerase can go on many laps around a library if we let it, and with each lap we become more confident about what the sequence is. After at least three times on the Sequel II, we can be confident that 99 out of 100 bases sequenced are correct.”
Barker, who is enjoying his tenth year at the Earlham Institute, explains that the technologies have changed rapidly and continuously during his time here.
“Illumina sequencers initially worked by using a laser to excite a different fluorophore for each base, which would give off a flash of light captured by a camera,” he tells us. “Now most of them use two fluorophores, whereby two of the bases have their own fluorophore, one has a combination of both, and the other doesn’t have a fluorophore at all.
“It basically takes two images, one for the fluorophore excited by the red laser and one for the fluorophore excited by the green, and we can tell what DNA bases are present.
“Pacific Biosciences uses fluorophores as well but it’s a bit different, whereby the polymerase is anchored at the bottom of a well they call a ‘zero mode waveguide’, the bottom of which is illuminated with a laser. All four bases have their own fluorophore, and when one of them binds to the DNA it stays there for slightly longer, allowing us to distinguish it from the other bases passing through the bottom of the well.”
“Because it uses tiny nano lenses positioned under each well to capture the information, it’s as though we’re filming each well,” adds Irish. “It's why Pacific Biosciences calls the data acquisition a ‘movie’.”

Assembling the data
Armed with the best possible sequence data, senior computational biologist Dr Graham Etherington of the Haerty Group could then get to work on assembling the tilapia genome.
“I looked at various different ways of assembling the data,” says Etherington, reflecting on the process. “I first used the Hi-Fi data to assemble the genome, and then used the extra consensus long-read information to scaffold it further and fill in any gaps. We’ve then got the 10X data on top.”
A scaffold, in genome assembly, contains complete DNA sequences in a general order and orientation that we can be relatively confident is correct. Scaffolds will have gaps, and we can perhaps imagine the scaffold to be somewhat like its namesake in construction, maintaining the structure of the genome while we fill in the gaps with the proper building blocks.
To get to the final assembly, Etherington used a number of tools.
“There’s a great tool called Tigmint,” he explains. “It aligns reads produced from the same molecule of DNA to your assembly and then breaks it in the parts where the reads don’t match. It also sticks scaffolds together that weren’t joined in the original assembly, but have matching sequences.
“Ross Houston at the Roslin Institute provided us with a linkage map produced by crossing 1,113 GIFT tilapia, allowing us to order and orientate almost 95% of our assembly into chromosome-scale sequences.”
Annotations
Having a well-constructed genome is just the beginning when it comes to using the information. It’s also important to annotate it - to place the crucial genes and other genetic elements that we can target in breeding programmes.
For that, Postdoctoral Researcher Dr Will Nash worked closely with senior software developer Luis Yanes and senior bioinformatician and developer Gemy Kaithakottil of the Swarbreck Group, who have developed the software applications to do just that.
“The first part,” says Nash, “is using the annotations of reference species O. niloticus and O. mossambicus, which are the species that have contributed to the genetic makeup of the GIFT strain.”
Because we already had a reference genome for these fish, it was possible to map any gene models we already knew about to our new GIFT assembly. Nash used a liftover approach combining the Liftoff software and the in-house ei-liftover pipeline generated by the Swarbreck Group that gives us an idea of how many of those accurately match our new genome.
“Ei-liftover compares the gene models that are predicted to be in the target species with the gene models present in the original, and looks at how accurate they are,” Nash explains. “It allows you to filter the results, leaving gene models that match 100% to the new assembly.”
Next, Nash used another Swarbreck Group pipeline called ‘REAT transcriptome’ to match gene expression, or RNA-seq, data from the closely related reference species to the new GIFT assembly.
“We found the mapping was almost identical,” says Nash. “So we could use RNA-seq from the two ‘parent’ species to describe where the genes were in GIFT.”
The process can be mirrored with protein sequences, this time using the ‘REAT homology’ tool.
“Essentially we’re protecting gene models at all three levels, RNA, DNA and protein, to give us the most accurate picture,” says Nash. “When we put all of these together using another tool from the Swarbreck group; Minos, and in our draft annotation, we get about 45,000 protein coding genes projected, with 31,000 classified as high confidence.
“Using BUSCO, a tool we use to assess the completeness of a genome assembly or an annotation by searching for single copy orthologs, we get a score of 99% of the BUSCO ortholog set based on protein similarity. So it seems like we have a very complete annotation.”

I looked at various different ways of assembling the data, I first used the Hi-Fi data to assemble the genome, and then used the extra consensus long-read information to scaffold it further and fill in any gaps. We’ve then got the 10X data on top

Dr Graham Etherington, Senior Computational Biologist, Earlham Institute

The holy grail of DNA sequencing
All of this means that we now have a very complete genome with which collaborators at WorldFish can breed more resilient fish.
“We’ve used the latest recipe at our disposal for these high quality genomes,” says Head of Genomics Pipelines Dr Karim Gharbi, whose team is now in the process of sequencing another tilapia strain for WorldFish.
“It’s nice for us to be able to bring all of these platforms together to give us assemblies that are as good as possible, making sure we haven’t missed anything out - any of that dark matter.
“Having the genome in a smaller number of larger blocks up to chromosome level is pretty much the holy grail of genome sequencing. Then there is how accurate it is, and I think these high-quality Hi-Fi reads we have generated are really making a difference.
“We’ve got genomes that are as representative of the real thing as currently possible.”
With some further work to be announced in the future, the genome assembly - which has already been used to boost tilapia breeding in Zambia - will benefit from whole chromosome length mapping.
“Because of the technologies applied, we should be able to further improve this assembly to chromosome length scaffolds and provide an assembly of even better quality than the current reference,” says Evolutionary Genomics Group Leader, Dr Wilfried Haerty.
“It's very exciting, because it opens up a lot of possible work with WorldFish on the future of these tilapia strains.”

We’ve used the latest recipe at our disposal for these high quality genomes, it’s nice for us to be able to bring all of these platforms together to give us assemblies that are as good as possible, making sure we haven’t missed anything out - any of that dark matter

Dr Karim Gharbi, Head of Genomics Pipelines, Earlham Institute
The research and development described in this article was funded by our UKRI-BBSRC Core Strategic Programme and National Capability in Genomics and Single Cell Analysis, in collaboration with partners at WorldFish.
If you would like to work with our Genomics Pipelines team on a sequencing project or a research collaboration, visit our Genomics Services page.
-
1 - Genome annotation workflow incorporating repeats annotation, RNA-Seq assemblies, Portcullis RNA-Seq junctions, protein alignments and utilising Mikado, for both proteins and transcripts, as a way filtering the data. AUGUSTUS gene builds were generated using different evidence inputs or weightings and were supplemented with gene models derived from protein alignments and gene models from the Mikado transcript selection stage. The Minos-Mikado pipeline was run to generate a consolidated set of gene models, the pipeline utilizes external metrics to assess how well supported each gene model is by available evidence, based on these and intrinsic characteristics of the gene models a final set of models was selected.