NanoOK of the North and South: A story of life in the Antarctic
Technology and Algorithms Group Leader Dr Richard Leggett tells us about his pioneering software tools NanoOK & NanoOK RT, which are making an impact in life sciences research.
No, we're not talking about the Inuit "master of the bears". Here, in fact, our Group Leader Dr Richard Leggett tells us about how his pioneering software NanoOK is making an impact in life sciences research.
1. The original NanoOK allows alignment-based analysis of nanopore sequencing data. NanoOK will automatically generate a PDF containing comprehensive analysis of data quality, accuracy and error profile.
2. You can also compare different samples and NanoOK will generate a comparison report.
3. A new version, NanoOK RT, provides real-time analysis of metagenomic samples, while sequencing progresses. It will also report on antibiotic resistance profiles.
The NanoOK and NanoOK RT software applies to the latest technology in Nanopore sequencing which enables scientists to retrieve live data through the MinION - a handheld DNA-sequencing device developed by Oxford Nanopore. The innovative device has opened up ‘in-the-field’ sequencing technology for human health and environmental projects.
The MinION works by detecting individual DNA bases that pass through a nanopore, and unlike existing sequencing technologies, there are few inherent sensing limits on the length of the DNA sequence that it can read in one go.

Nanopore sequencing is a relatively new field and new tools are needed to get the most out of the technology. That’s where NanoOK and NanoOK RT come in.
At EI, we’ve used it to trial nanopore sequencing as a diagnostic tool for preterm babies suffering intestinal diseases. We recently used it to carry out in-field analysis of extreme microbiome samples in Iceland and later in the year, we’ll be using it on board a research vessel in the Antarctic.
Ship-Seq: Nanopore sequencing of polar microbes onboard Icebreakers
The Antarctic is a core barometer for how climate change is impacting our planet. PhD Student Emma Langan in the Leggett Group is currently on expedition in Antarctica, using the Nanopore MinION onboard Icebreaker ships to live sequence polar phytoplankton and other microbes to explore the effects of global warming.
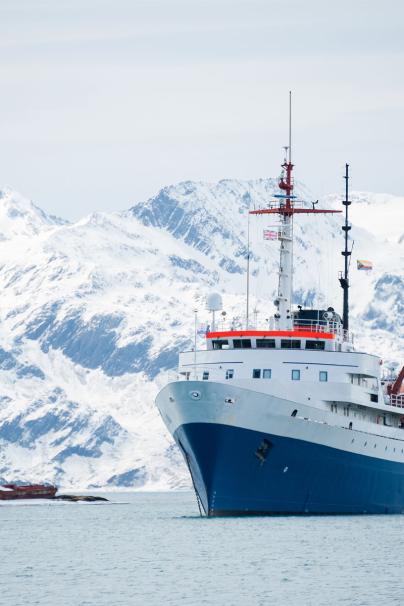

A proof-of-concept study for using the MinION on research vessels in-situ to prevent loss of information from long-term storage of samples and to allow decisions to be made in real-time based on present species.
"Emma Langan will be undertaking fieldwork in the Southern Ocean with the British Antarctic Survey on the RRS Discovery. Metagenomic sequencing of polar ocean water samples will be performed using the Nanopore MinION."
"While the sequencing is ongoing, the NanoOK reporter will provide live information on the living organisms in the sample. This experiment is to determine whether MinION sequencing can be used effectively in-situ and the practicality of real-time reporting. On-board sequencing will be compared to lab-based sequencing back in the UK."
The potential benefits of this approach are manifold;
a) real-time assessment of in-situ microbial diversity
b) real-time analysis of in-situ or onboard experiments in polar oceans and
c) genome and transcriptome sequencing of sensitive but ecologically relevant polar microbes.
“If successful this will be trialled in other polar ocean cruises, focusing more on high molecular weight DNA sequencing so that we can try to build a database of reference assemblies for polar microbe species. Currently there are very few available which makes investigating genetic adaptations to changing climate conditions difficult.”
The tool was published in the journal Bioinformatics and is well used within the community and cited in a wide range of publications.
I’m most pleased with the pre-term baby diagnostics work. This was what we originally developed NanoOK RT for.
Whilst we’ve only done a proof-of-concept so far, rolling this out as an aid to clinicians in a real setting has the potential to make a real impact on treatment and outcomes.
The original NanoOK’s alignment-based analysis of nanopore data isn’t really reproduced by other software.
There is other software that aims to do a similar analysis to NanoOK RT, but it’s commercial software that requires payment for each analysis, isn’t extensible and requires a fast internet connection.
The latter makes the commercial software difficult for many in-field applications.
I’m most excited by in-field applications (negating the logistical problems associated with sample transport) or applications that are particularly time sensitive (e.g. pathogen identification).
These are the ones where NanoOK RT can make the biggest difference.
The group is currently working on producing a new version of NanoOK RT that provides a nicer web interface and more sophisticated analysis. Look out for that in the next year.
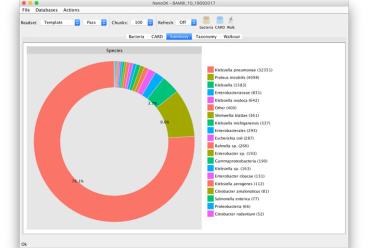
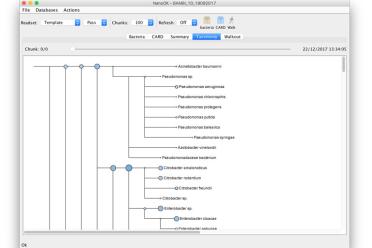
NanoOK is a tool for comprehensive alignment-based analysis of nanopore sequencing data.
NanoOK RT extends NanoOK to provide functionality for real-time analysis and classification of metagenomic samples.