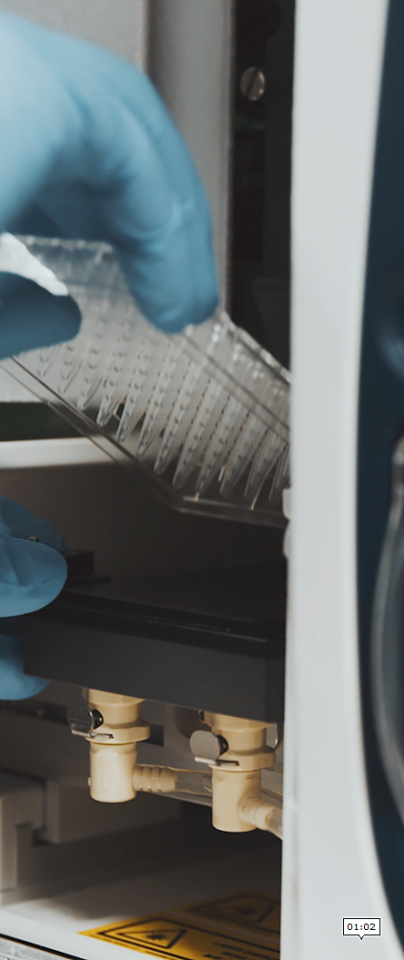
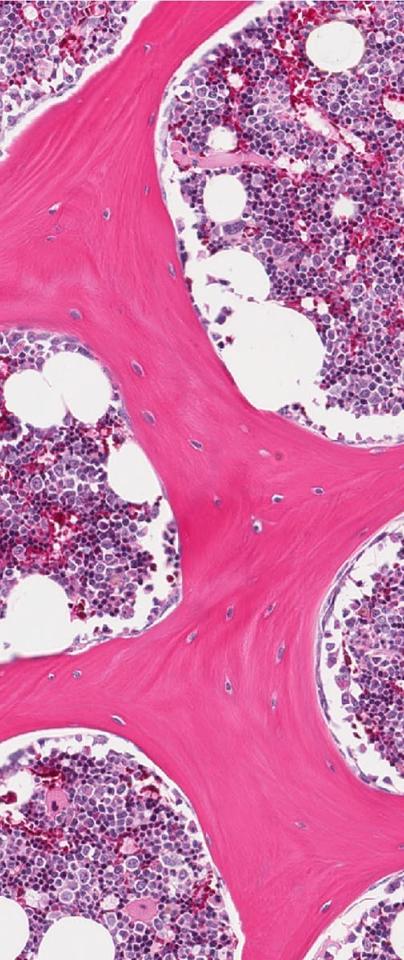
Though we know that stem cells in the bone marrow are able to produce offspring that lead to all of the cell types we see in the blood - red, white, and platelets among them - how they get there is hard to determine.
“My work is heavily focused on megakaryocytes - the cells that break into the fragments we call platelets,” says Scoones, describing her PhD project. “It was thought that these megakaryocytes arose from a bipotent progenitor, which means, basically, one cell that could generate either megakaryocytes or red blood cells.”
As Scoones has seen first-hand through her work, it’s not so (relatively) simple.
“What we've seen is that, to generate a megakaryocyte, you can actually go straight from a stem cell to a megakaryocyte progenitor, without these stepwise transitions. There's some kind of bias in the stem cells that primes them for platelet-specific gene expression, but the origin of that hasn’t been fully explored yet.”
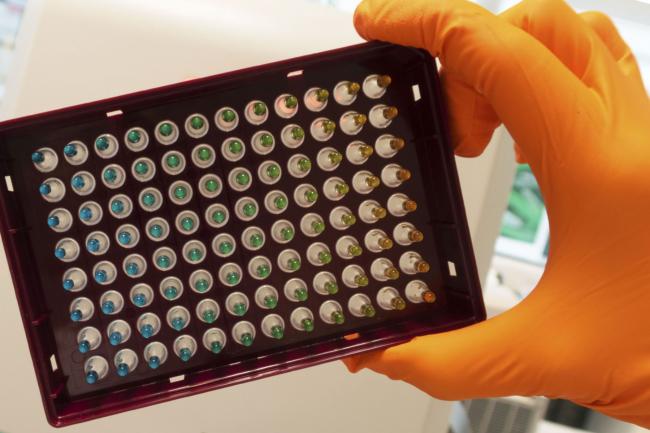
Within a sample of cells that Scoones is able to extract from bone marrow, it’s possible to identify stem cells based on markers amongst the genes they express. Some of those cells will be biased towards producing immune cells, red blood cells, or indeed megakaryocytes. The mechanisms that prod them in that direction, however, still elude us.
Perhaps there are triggers that might persuade a cell along a certain direction.
“What's really interesting is that there seem to be cell behaviours that can be activated through stress,” says Scoones. “We think that by triggering an 'emergency' pathway, such as low platelet levels, we might discover the key genes driving stem cell differentiation, which is something I’m focusing on at the moment.