At EI we tend to work on things that don't have a protocol yet. We do lots of technology development, which is really fun.
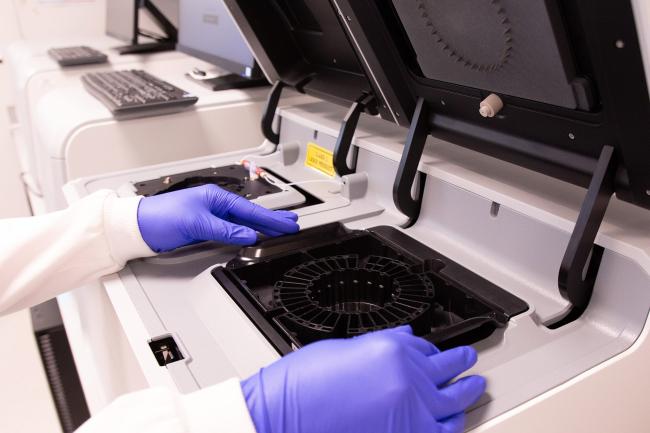
We have an unbelievable suite of fancy instruments to do that - it’s outstanding. People approach us to look at really weird and wonderful things, which means that we have several different instruments for single-cell isolation that are all excellent for different applications. We essentially have a cell-sorting suite, which is quite unique to the Institute and an asset to the region.
On top of that, we have robotics and automation platforms - and we are lucky to be working directly with the sequencing team in Genomics Pipelines. That really helps because I've worked in that group, so I am constantly talking to them and asking them for advice and vice versa.
I can receive a sample and have it sequenced within the same week, potentially. That's all because everything is in the same building. We have the expertise for the whole pipeline, really.
The other thing is that it’s very unusual to have a boss [Iain Macaulay] who knows how to run all of the instruments, and how to do this stuff in the lab at the ground level. That's very, very useful. He knows that things sometimes go awry, and he has advice and backup plans as well. That's also very good for setting up collaborations, as he can be in those meetings and understand projects from the outset.