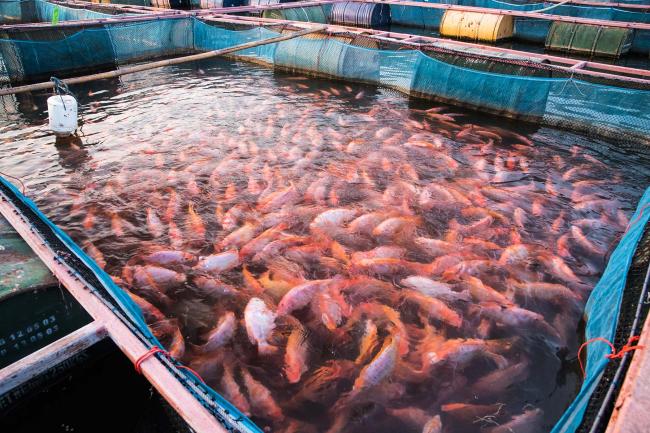
I am a Postdoctoral Research Scientist in the Haerty Group, working on the Tilapia Genomic Resources project. As part of this project, I will be using bioinformatic and transcriptomic techniques including bulk and single-cell RNA-Seq. My work aims to identify loci associated with aquaculture relevant traits related to environmental stress, including temperature, salinity and pathogen resistance within Tilapia species.
Prior to this position, I was a NERC ARIES DTP PhD student at the University of East Anglia, the Natural History Museum London and Chester Zoo working on the conservation genomics of endangered bird species in zoos.
This project focused on quantifying the genetic load of individuals in captive breeding programs using CADD scores within the ultraconserved elements of the genome. These methods were applied to species that had been through intense population bottlenecks including the pink pigeon (Nesoenas mayeri) and the whooping crane (Grus americana). These approaches were designed to inform optimal mate pairings for captive breeding and reintroduction programs, to avoid inbreeding depression.
In 2019 I completed my BSc Hons in Biological Sciences at the UEA and in 2020 my MScR where I applied bioinformatic analyses to detect signature of genetic drift and adaptive evolution in diatoms at the UEA.